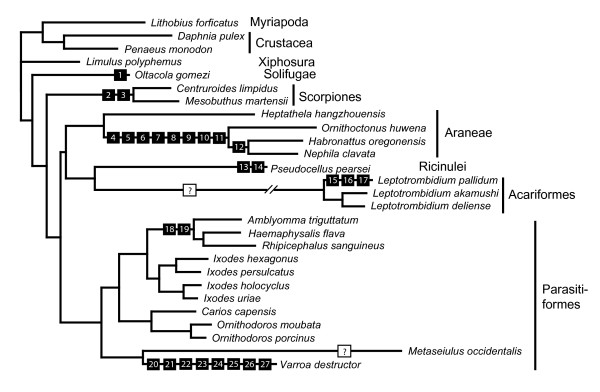
Figure 6.
Hypothesized events leading to derived gene orders (black squares, compare Fig. 5) and strand bias reversal (red circles, compare Table 2) mapped on a topology based on the results of our phylogenetic analysis (compare Fig. 4, amino acid data set). 1: Translocation of trnP; 2: Reversal in nucleotide bias; 3: Translocation of trnQ; 4: Reversal in nucleotide bias; 5: Translocation of trnI; 6: Translocation of trnQ; 7: Translocation of trnY or trnC; 8: Translocation of trnL; 9: Translocation of trnN; 10: Translocation of trnS; 11: Translocation of trnT; 12: Translocation of trnI; 13: Translocation of trnN; 14: Tandem duplication of a gene block ranging from trnL to trnY followed by random deletion; 15, 16: Two subsequent tandem duplications of a gene block ranging from trnQ to the large non-coding region followed by random deletion; 17: Duplication and inversion of rrnL and a large non-coding region, probably by nonhomologous intergenome recombination (15–17 according to [59]); 18: Translocation of trnC; 19: Tandem duplication of a gene block ranging from trnF to trnQ followed by random deletion; 20: Reversal in nucleotide bias; 21: Tandem duplication involving the putative control region and rrnS followed by random deletion; 22: Translocation of trnH; 23: Translocation of trnQ; 24: Translocation of trnY; 25: Translocation of trnP; 26: Translocation of trnS; 27: Translocation of trnC; "?": Probably due to multiple rearrangements, the history of events leading to the highly derived gene order of Leptotrombidium and Metaseiulus cannot reliably be reconstructed.