Figure 3.
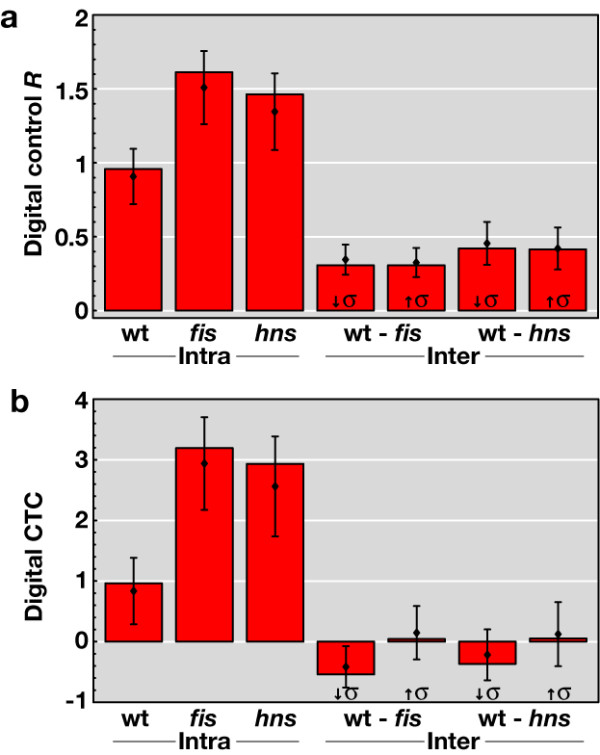
Digital control R and control type confidence (CTC) of transcriptional regulation. (a) The digital control R is the number of connected nodes divided by the number of isolated nodes in the effective transcriptional regulatory networks of the three intra-strain experiments (wt, fis, hns) and the four inter-strain experiments (wt-fis and wt-hns for low (↓σ) and high (↑σ) negative supercoling each). (b) Digital CTC quantifies the deviation of the effective subnet based on significant expression changes from an appropriate null model. To estimate the sensitivity of the observables against noise, we replace 10% of all affected genes with randomly selected genes from the pool of affected genes in all other experiments. We then recalculate the digital control via the ratio R (a) and the corresponding CTC (b). We show the mean Rs (diamonds) together with the standard deviation for 10,000 runs, and the mean CTCs (diamonds) together with the standard deviation for 1,000 runs, where the actual data is compared to 1,000 null model runs each.
