Fig. 4. Specific histone modifications induced at hypoxia-responsive genes.
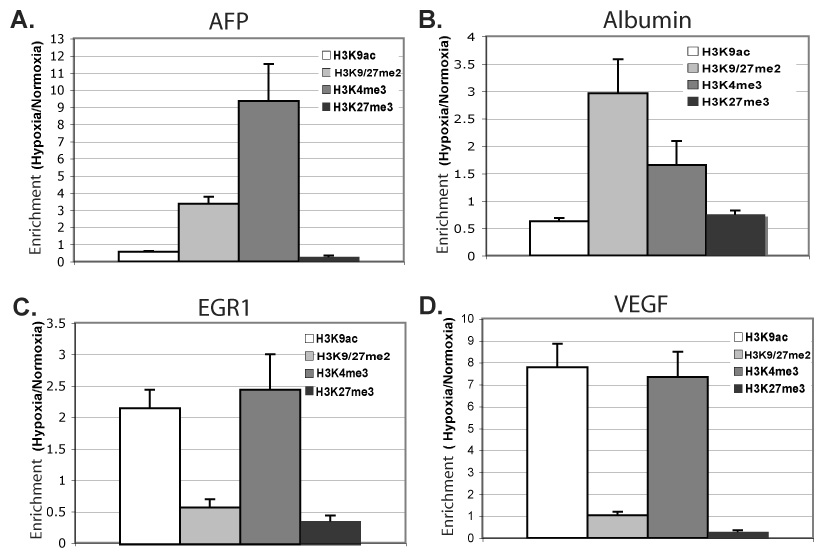
ChIP of indicated modified histones in Hepa 1–6 cells after 48h at normoxia or 0.2% oxygen, followed by quantitative PCR for core promoter regions of AFP (A), Albumin (B), EGR1 (C), and VEGF (D). The %-input bound by each modified histone H3, minus background (IgG), was normalized by total bound histone H3; fold-change hypoxia versus normoxia is graphed. Averages represent of two to three independent experiments, each experiment quantified in duplicate, with error bars representing SEM.
