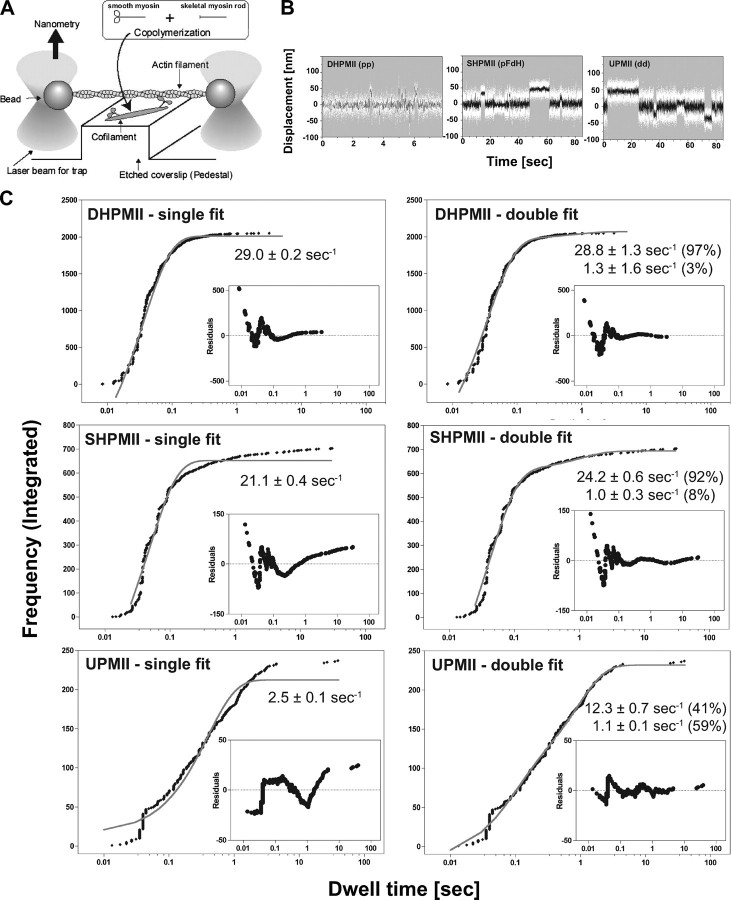
FIGURE 2.
Single molecule mechanical measurement of smooth muscle myosin with optical trap nanometry. A, the nanometry setup for measuring actin displacements by a single smooth muscle myosin molecule. B, typical recordings of actin displacements by DHPMII, SHPMII, and UPMII. The white dots represent raw data. The data passed through a low pass filter of 30-Hz bandwidth is shown by black lines. Note the differences in the time scale among the recordings. C, dwell time distribution of the actomyosin interaction. The integrated frequencies of DHPMII (pp) (n = 2051), SHPMII (pFdH) (n = 704), and UPMII (dd) (n = 237) were plotted against the observed dwell times. Both single exponential and double exponential fitting were tested, and the residuals are shown in the inset graphs. Note that the differences in the amplitude of the residuals among the smooth muscle myosin species came from their different sampling sizes (n = 2051, 704, and 237 for DHPMII, SHPMII, and UPMII, respectively). The single exponential fitting gave apparent rates of 29.0 ± 0.2, 21.1 ± 0.4, and 2.5 ± 0.1 s–1 for DHPMII, SHPMII, and UPMII, respectively, whereas the double exponential fittings gave 28.8 ± 1.3 s–1 (97%) and 1.3 ± 1.6 s–1 (3%) for DHPMII, 24.2 ± 0.6 s–1 (92%) and 1.0 ± 0.3 s–1 (8%) for SHPMII, and 12.3 ± 0.7 s–1 (41%) and 1.1 ± 0.1 s–1 (59%) for UPMII. The percentage values are the amplitude contributions of the different rate components. The amplitudes of the fitting curves were greater than the total integrated frequencies because displacements with very short dwell times were not detected because of the 30-Hz low pass filtering of the raw data. (The shortest possible events we can detect with the 30-Hz low pass filter can be calculated as 1/2 Pi/30 =∼5 ms. Thus the dead time of our measurement is ∼5 ms.) Note that the dwell time distribution of DHPMII is fairly explained by a single exponential kinetics, whereas those of SHPMII and UPMII are well explained by double exponential.