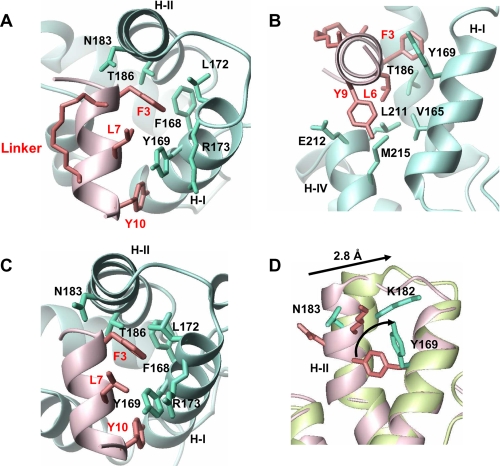
FIGURE 2.
Structural details of intermolecular contacts with ribbon representation of the protein (blue) and peptide (pink) backbone. A, the top view of the binding surface displays the interactions between the side chains of Phe-3 and Tyr-10 from the peptide and helix I and II of mCA-CTD. B, the side view of the complex displays the interactions that anchor Leu-6 and Tyr-9 from the peptide using Leu-211 and Met-215 from helix IV. C, the top view of the x-ray structure of CAI in complex with CA-CTD (2BUO). D, superposition of the backbone Cα atoms of CA-CTD (pink) and mCA-CTD (green) based on alignment generated from residues in helix I, helix III, and helix IV (r.m.s.d. = 0.8 Å). When helix II is included, the r.m.s.d. increases to 1.3 Å. Residues that are important for binding the target peptide and rearranged through the helix movement are indicated in the figure. The PDB code for CA-CTD structure used in the alignment is 1A8O. The figures were generated in MOLMOL 2.1 (30).