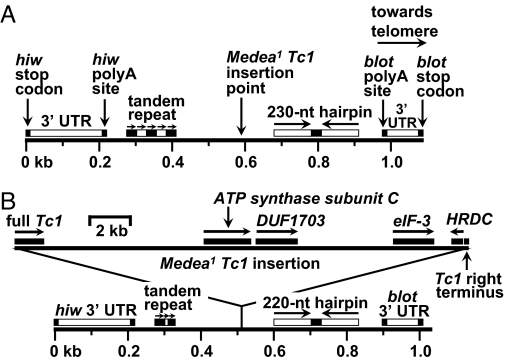
Fig. 2.
The M1-associated insertion is a 21.5-kb composite Tc1 element. (A) Map of hiw-to-blot region of a wild-type chromosome. The two genes (tail-to-tail orientation) are separated by a span of 1.1 kb (stop codon to stop codon). Vertical arrows indicate stop codons, polyA addition sites, and Tc1 insertion site. The insertion point is flanked by five copies of a 27-nt, imperfect tandem direct repeat, as well as a 230-nt hairpin consisting of a 20-nt central loop flanked by a pair of 105-nt, imperfectly palindromic stems. Horizontal arrows indicate the tandem direct repeats, hairpin stems, and orientation of the entire segment on the chromosome. Horizontal bars above insertion target chromosome denote main features of region flanking the M1 insertion target. (B) Map of hiw-to-blot region of an M1 chromosome. Solid bars above the 21.5-kb composite Tc1 insertion (top part of B) denote genes or gene fragments within the insertion. Horizontal arrows indicate directions of transcription. Of these genes (Tc1 transposase, ATP synthase subunit C, DUF1703, eIF-3, and HRDC) only DUF1703 is thought to be potentially functional (see Fig. S3). The isolated Tc1 right terminus is not shown to scale. Note that the tandem repeat and hairpin are shorter in the M1 strain than in wild type (compare A and B). Also note the different scales for insertion vs. flanking regions.