Table 2. Structure-refinement statistics.
| Resolution range | 23–1.95 |
| No. of protein atoms | 2572 |
| No. of cadmium/chloride ions | 1/1 |
| No. of water molecules | 226 |
| Rcryst† (%) (No. of reflections) | 21.0 (27186) |
| Rfree† (%) (No. of reflections) | 25.7 (1486) |
| R.m.s.d. bonds (Å) | 0.020 |
| R.m.s.d. angles (°) | 2.16 |
| Average B factor (Å2) | 25.3 |
| Residues not modelled | Monomer A/B, tag residues, 177–183, 1–3; monomer A, 4, 15–18, 126–129 |
| Residues in multiple conformations | B123, B138 |
| Ramachandran plot, nonglycine/proline residues | |
| In most favoured regions | 267 (97.1%) |
| In additionally allowed regions | 8 (2.9%) |
R
cryst = 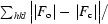
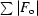 , where F
o is the observed structure-factor amplitude and F
c is the calculated structure-factor amplitude. R
free is calculated based on 5% of reflections not used in refinement.
, where F
o is the observed structure-factor amplitude and F
c is the calculated structure-factor amplitude. R
free is calculated based on 5% of reflections not used in refinement.
