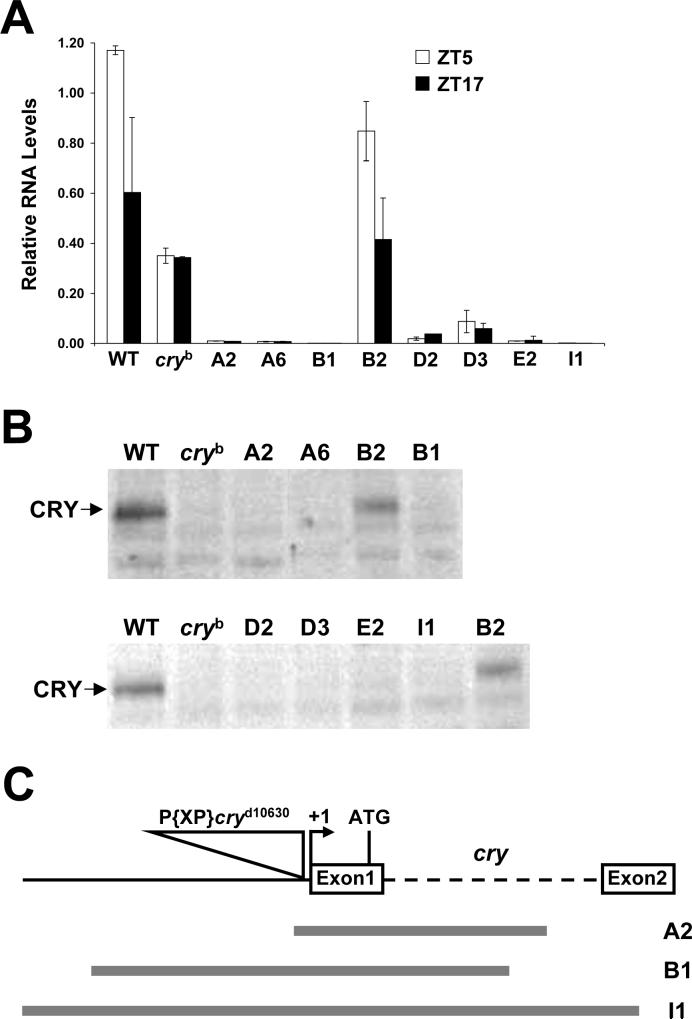
Figure 2.
Expression and structure of cry deletion mutants. A. qPCR of cry mRNA from the heads of w (WT), cryb and cry excision flies (A1, A6, B1, B2, D2, D3, E2, I1) collected at ZT5 (white bars) or ZT17 (black bars) under LD conditions. Relative cry mRNA levels were quantified as described in Materials and Methods. Error bars are ±SD. B. Western blots of head extracts from w (WT), cryb and excision flies (A1, A6, B1, B2, D2, D3, E2, I1) collected at CT19 during the first day of DD were probed with GP23. In panels A and B, B2 is a precise excision control. C. Structural characterization of cry deletion mutants. Diagram of the cry gene (top) showing the first two exons (white boxes), intron 1 (dashed line), upstream sequence (black line), the P{XP}cryd10630 insert (triangle), the transcription start site (+1) and the translation start site (ATG). The extent of deleted sequences the A2, B1 and I1 strains are shown as gray lines.