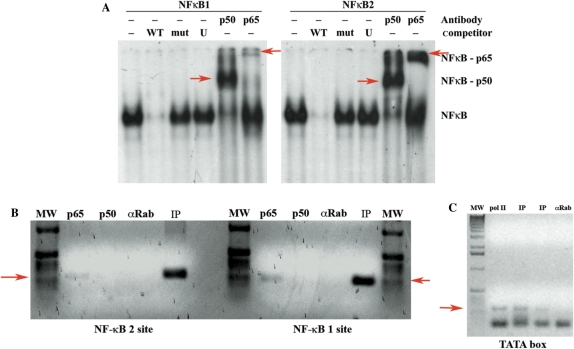
Figure 4.
The two putative NF-κB binding sites are bound by NF-κB complexes in vitro and in vivo in MC3 cells. (A) Electrophoretic mobility shift assay using nuclear extracts from MC3 cells (EBV-positive cells) on the two NF-κB binding sites. Left panel: NF-κB site 1 (−1150). Right panel: NF-κB site 2 (−1697). WT = 80-fold molar cold wild-type competitor; mut = 80-fold molar cold mutant competitor; U = 80-fold molar cold unspecific competitor; p50 = supershift with anti NF-κB p50 antibody; p65 = supershift with anti NF-κB p65 antibody. Supershift bands are indicated with red arrows. (B) ChIP assay of chromatin extracted from MC3 cells using polyclonal antibodies directed against the NF-κB p65 or NF-κB p50. After immunoprecipitation, the DNA samples were purified and subjected to PCR with specific primers amplifying the NF-κB site 1 region (right panel) or NF-κB site 2 region (left panel). MW = molecular weight marker; p65 = pAB anti NF-κB p65; p50 = pAb anti NF-κB p50; αRab = unspecific polyclonal antibody anti rabbit immunoglobulin; IP = input. The red arrows indicate the specific amplified band. (C) ChIP assay of chromatin extracted from MC3 cells using monoclonal antibody directed against the RNA pol II. After immunoprecipitation, the DNA samples were purified and subjected to PCR with specific primers amplifying the TATA box region. MW = molecular weight marker; pol II = mAB anti RNA pol II; IP = input. The two lanes represent two different amounts of input sample used for the PCR; αRab = unspecific polyclonal antibody anti rabbit immunoglobulin. The red arrow indicates the specific amplified band.