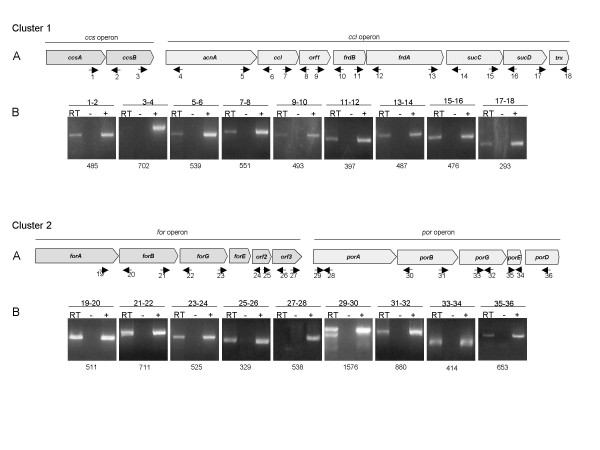
Figure 2.
Structure and genetic organization of the L. ferriphilum DSM 17947 genes in cluster 1 and cluster 2 predicted to be involved in reductive tricarboxylic acid (RTCA) cycle. A. Schematic map of the RTCA locus. Genes located in Cluster 1: ccsA (citryl-CoA synthetase, subunit A), ccsB (citryl-CoA synthetase, subunit B), acnA (Aconitase A), ccl (citryl-CoA lyase), orf1, frdA (fumarate reductase, subunit A), fdrB (fumarate reductase, subunit B), sucC (succinyl-CoA synthetase, beta subunit), sucD (succinyl-CoA synthetase, alpha subunit) and trx (thioredoxin). Genes located in Cluster 2: forA (2-oxoglutarate ferredoxin oxidoreductase, alpha subunit), forB (2-oxoglutarate ferredoxin oxidoreductase, beta subunit), forG (2-oxoglutarate ferredoxin oxidoreductase, gamma subunit), forE (2-oxoglutarate ferredoxin, epsilon subunit), orf2 (hypothetical protein), orf3 (hypothetical protein), porA (pyruvate ferredoxin oxidoreductase, alpha subunit), porB (pyruvate ferredoxin oxidoreductase, beta subunit), porG (pyruvate ferredoxin oxidoreductase, gamma subunit), porE (pyruvate ferredoxin oxidoreductase, epsilon subunit), porD (pyruvate ferredoxin oxidoreductase, delta subunit). B. RT-PCR amplification (RT) of intergenic. -, RT-PCR amplification control without reverse transcriptase; +, standard PCR amplification control. Primer pairs used for amplification are indicated at the top of each panel and amplification product sizes are indicated at the bottom of each panel.