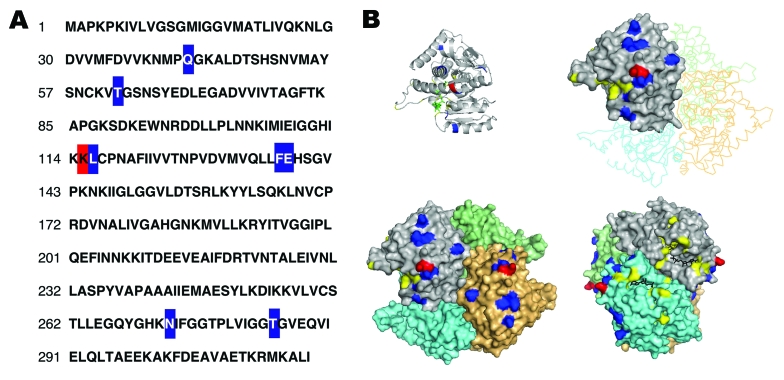
Figure 2.
Modeling of the analysis of Plasmodium knowlesi lactate dehydrogenase (LDH). A) Sequence of LDH from P. knowlesi deduced from genomic DNA fragments sequenced by the Sanger malaria genome project (www.sanger.ac.uk/Projects/P_knowlesi). LDH isoforms from P. vivax, P. malariae, P. ovale, P. berghei, P. yoelli, and P. falciparum were compared with that of P. knowlesi. Residues unique to P. knowlesi and P. vivax are shown in blue; residues unique to P. knowlesi and P. falciparum are shown in red. B) Model of P. knowlesi LDH and specific epitopes. A model for P. knowlesi LDH was calculated by using WURST protein threading server (www.zbh.uni-hamburg.de/wurst/index.php) and the P. falciparum and P. vivax crystal structures (PDB: 2A94 and 3 2AA3). Shown is the monomer, as well as the assembled tetramer, aligned to the backbone of the P. vivax tetramer using pymol. The nicotinamide adenine dinucleotide cofactor analog 3-acetyl pyridine adenine dinucleotide is shown in black. Residues important for substrate binding and catalysis are shown in yellow. P. knowlesi residues shared only with P. vivax are shown in blue and indicate where the 11D9/13H11 epitopes could be. P. knowlesi residues shared only with P. falciparum are shown in red and indicate a critical determinant of the 17E4/7G9 epitopes.