Figure 1.
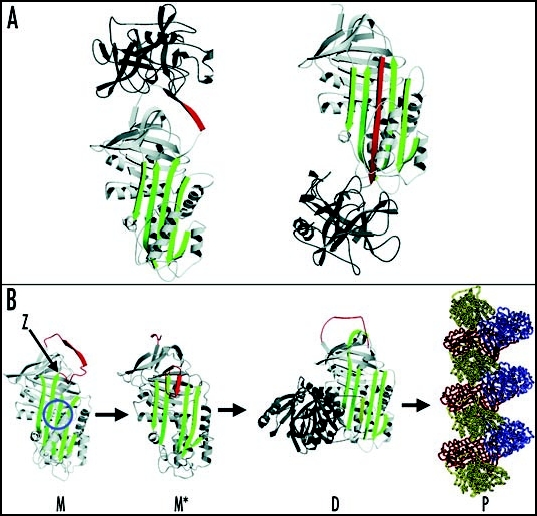
Inhibition of neutrophil elastase by α1-antitrypsin and the structural basis of polymerization. (A) After docking (left) the neutrophil elastase (grey) is inactivated by movement from the upper to the lower pole of the protein (right). This is associated with the insertion of the RCL (red) as an extra strand into β-sheet A (green). (B) The structure of α1-antitrypsin is centred on β-sheet A (green) and the mobile reactive centre loop (red). Polymer formation results from the Z variant of α1-antitrypsin (Glu342Lys at P17; indicated by arrow) or mutations in the shutter domain (blue circle) that open β-sheet A to favour partial loop insertion and the formation of an unstable intermediate (M*). The patent β-sheet A then accepts the loop of another molecule to form a dimer (D), which then extends into polymers (P). The individual molecules of α1-antitrypsin within the polymer, although identical, are coloured red, yellow and blue for clarity. Figure reproduced with permission from Lomas et al.97
