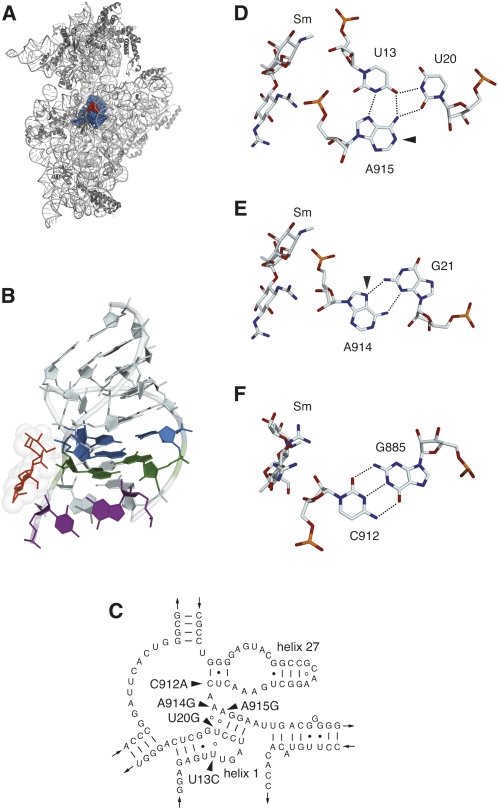
FIGURE 1.
(A) Location of the central pseudoknot (blue spheres) and the streptomycin (red) binding site in the T. thermophilus 30S subunit crystal structure (Carter et al. 2000). (B) Structure of the central pseudoknot, showing the U13–A915–U20 base triple (blue), the G21–A915 base pair (green), and the C912–G885 base pair (purple). Streptomycin is shown as red sticks with van der Waals radii indicated. (C) Secondary structure model of the central pseudoknot region of T. thermophilus HB8 16S rRNA (Cannone et al. 2002) modified to reflect the base pairing and base triple arrangements observed in the 30S subunit crystal structure (Wimberly et al. 2000). Sites of base substitutions conferring streptomycin resistance are indicated. (D) The U13–U20–A915 base triple and streptomycin (Sm), showing the hydrogen bonding network (dashed lines) and the site of DMS hyperreactivity (arrowhead) at the N1 of A915. (E) The G21–A914 type II sheared G–A base pair and streptomycin. The arrowhead indicates the N7 of A914 that becomes reactive to DMS in the A914G mutant. (F) The C912–G885 Watson–Crick pair and streptomycin. (A,B,D–F) Generated with PyMol (DeLano 2002) using PDB file 1FJG (Carter et al. 2000).