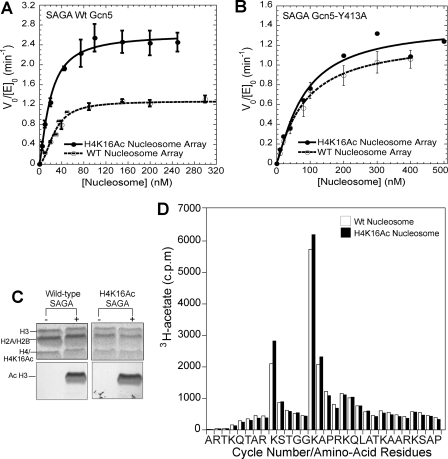
FIGURE 4.
H4-Lys-16 (H4K16Ac) acetylation promotes H3 tail acetylation turnover in a bromodomain-dependent manner. A, a comparison of HAT activity of wild-type SAGA complex on nucleosome arrays containing H4-Lys-16 acetylation (solid circles) and wild-type nucleosome arrays (open circles). From the fitting of the data, for H4-Lys-16 acetylated nucleosomal arrays (solid line), the half-saturation concentration is 19.7 ± 1.7 nm, and the kcat, app is 2.64 ± 0.29 min-1. For the arrays that were not preacetylated (dashed lines), the half-saturation concentration is 33.6 ± 5.0 nm with a kcat, app of 1.42 ± 0.15 min-1. B, a comparison of HAT activity of the SAGA Gcn5-Y413A complex on nucleosome arrays containing H4-Lys-16 acetylation (solid circles) and wild-type nucleosome arrays (open circles). Two independent experiments were performed for H4-Lys-16 acetylated nucleosomes at each nucleosome concentration. Fitting of the data give a half-saturation concentration and kcat, app of 98.1 nm and 1.50 min-1 for the H4-Lys-16 acetylated nucleosome arrays (solid line) and 104.7 ± 29.7 nm and 1.46 ± 0.15 min-1 for the wild-type nucleosome arrays (dashed line). C, a comparison of the histone specificity of the SAGA complex for wild-type (lane 2) and H4-Lys-16 acetylated (lane 4) nucleosome arrays. Histones are resolved by SDS-PAGE. Staining with Coomassie Blue is shown in the upper panel, with the corresponding fluorogram shown in the lower panel. D, a comparison of the sites of SAGA-mediated acetylation for nucleosomal arrays that are not preacetylated (white column) and for nucleosome arrays that are preacetylated at H4-Lys-16 (black column). The amount of [3H]acetyl incorporation at a given position of the H3 tail was determined by microsequencing. To more clearly compare the relative site selectivity between nucleosome substrates, the data for the H4-Lys-16 preacetylated substrate have been normalized so that its total H3 histone acetylation is equivalent to the total H3 acetylation for substrate that was not preacetylated.