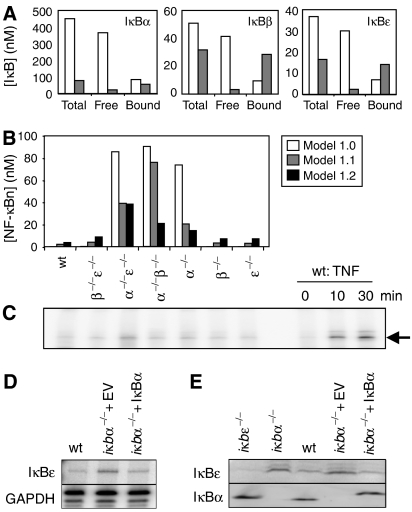
Figure 3.
An improved model of the homeostatic NF-κB signaling module. (A) Model calculations of IκBα, β, and ɛ protein levels (nM) in unstimulated cells for total IκB, free IκB, or NF-κB-bound IκB. Model 1.0 predictions are white bars, model 1.1 predictions are gray bars. (B) Model calculations of NF-κB activity (nM) in unstimulated wild-type and ikb−/− cells predicted by model version 1.0 (white bars), version 1.1 (gray bars), and version 1.2 (black bars). (C) NF-κB activity in untreated cells as measured by EMSA of nuclear extracts from the iκb−/− cell genotype labeled above each lane. The last three lanes are controls for the NF-κB band and are nuclear extracts from wild-type cells treated with TNF (1 ng/ml). (D) RNase protection assay showing levels of IκBɛ mRNA in untreated wild-type cells, iκbα−/− cells with empty vector control, or iκbα−/− cells expressing a retroviral iκbα transgene. GAPDH is used as a loading control. (E) Western blots for IκBɛ and IκBα in resting cells. The cell genotype is listed above each lane.