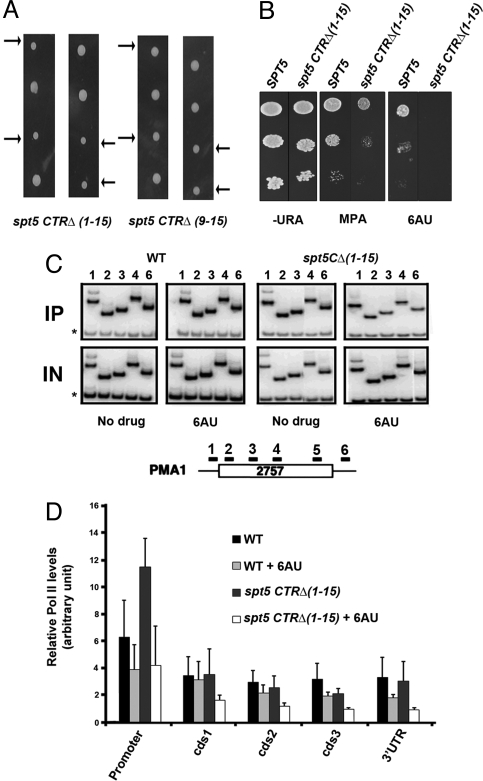
Fig. 2.
The Spt5 CTR is important for growth and promotes transcription elongation. (A) A diploid strain was transformed with a 3xHA-His3MX6 cassette replacing either the entire C-terminal region (repeats 1–15) of Spt5 or only part of the CTR (repeats 10–15). After sporulation, tetrads were dissected and individual spores were tested for growth. Left shows 2 representative tetrads from diploids in which all 15 repeats were removed. Right shows 2 representative tetrads from diploids in which 6 of 15 repeats were removed. The arrows point to the 2 spores (meiotic products) of each tetrad that segregate with the CTR deletion. (B) Deletion of the Spt5 CTR causes sensitivity to elongation inhibitors. Ten-fold serial dilutions of SPT5 and spt5ctrΔ(1–15) strains were tested for growth on plates in the presence of mycophenolic acid (MPA) or 6-AU. (C) Representative chromatin immunoprecipitation (ChIP) showing that elongation by RNAPII is less efficient when the Spt5 CTR is deleted. Chromatin was prepared from SPT5 and spt5 CTRΔ(1–15) strains grown in the presence or absence of 6-AU, as indicated. Chromatin was precipitated with 8WG16 monoclonal antibody that recognizes RNAPII and analyzed using primers specific for the PMA1 locus. A diagram of the PMA1 locus is shown to indicate the location of the ChIP primer pairs (from ref. 56). The asterisks indicate a nontranscribed sequence near the telomere of chromosome 5 that is coamplified in each PCR as a control for nonspecific precipitation. IN, input DNA; IP, precipitated DNA. (D) Quantification of ChIP data like that shown in Fig. 2C, with error bars indicating standard deviations. Relative enrichment was calculated as [(Experimental band IP)/(Control band IP)]/[(Input Experimental band)/(Input Control band)]. Promoter, cds1,2,3, and 3′UTR correspond to primer pairs 1, 2, 3, 4, and 6, respectively.