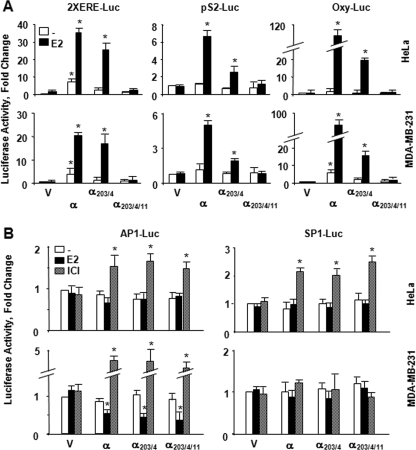
FIGURE 2.
Transcriptional responses to ERα proteins from reporter systems emulating the ERE-dependent and ERE-independent signaling pathways. A, HeLa or MDA-MB-231 cells were transiently transfected with an expression vector bearing none (V) or an ERα cDNA together with a reporter vector bearing two consensus ERE sequences in tandem located at the upstream of the simple TATA box promoter (2XERE-Luc), the proximal promoter from the trefoil factor 1, pS2 (pS2-Luc), or oxytocin (Oxy-Luc) gene in the absence or presence of 10-9 m E2 for 24 h. Simulating the ERE-dependent signaling pathway, these promoters drive the expression of the Firefly luciferase cDNA as the reporter enzyme. Cells were also co-transfected with a reporter vector bearing the Renilla luciferase cDNA for transfection efficiency. Normalized luciferase values are represented as fold changes compared with the luciferase values obtained with control vector in the absence of E2, which was set to 1. Shown are the means ± S.E. of three independent experiments performed in duplicate. B, effects of ligands on transregulatory activities of ERs in simulated ERE-independent signaling pathways. HeLa or MDA-MB-231 cells were transiently transfected with an expression vector bearing none (V) or an ERα cDNA together with reporter vector bearing an AP1 (AP1-Luc) or SP1 (SP1-Luc) promoter driving the expression of the firefly luciferase enzyme cDNA as the reporter. Cells were also co-transfected with a reporter vector bearing the Renilla luciferase cDNA for transfection efficiency. Cells were then treated with fresh medium without or with 10-9 m E2 or 10-7 m ICI for 40 h. Normalized luciferase values are represented as fold change compared with the luciferase values obtained without ligand, which was set to 1. Results are the means ± S.E. of three independent experiments performed in triplicate. Asterisk indicates significant difference.