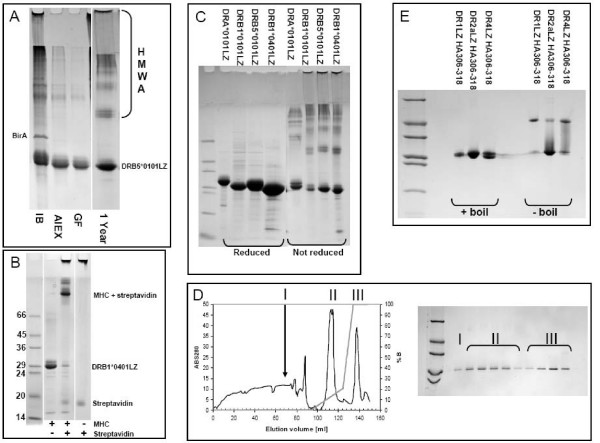
Figure 1.
Purification of MHC II and refolding of complexes. (A) Non reducing gel showing purification of urea denatured DRB5*0101. IB: urea denatured inclusion bodies after fermentation. AIEX: Purity after ion-exchange on Q-Sepharose FastFlow. GF: Purity after gel filtration on Superdex200, this is the final product from the purification. Though pure, the MHC band seems to consist of several sub bands each representing an oxidative isomer of the molecule. 1 Year: Same product after 1 year at -20°C. Note the formation of higher molecular weight aggregates (HMWA), and a more concentrated MHC band. (B) Biotinylation test, gel shift assay of DRB1*0401. Without streptavidin MHCbiotin travels as a monomer, addition of an excess of streptavidin causes nearly all MHCbiotin to shift to a position corresponding to a 1:1 complex with streptavidin, without MHCbiotin streptavidin travels as a monomer. In most cases a nearly quantitative biotinylation could be achieved. (C) Reducing and non-reducing SDS-PAGE of DRA*0101, DRB1*0101, DRB5*0101 and DRB1*0401. (D) IMAC purification of DR2a refolded in the presence of a histidine tagged HA306-318H6 peptide and eluted by an imidazole gradient. Fractions were collected and analysed by reducing SDS-PAGE (samples boiled). Peak I: Run through Peak II: empty α and β chain binding with moderate affinity to Ni2+ column Peak III: Elution of MHC II complex by imidazole. (E) Non-reducing SDS-PAGE analysis of DR1, DR2a and DR4 complexes refolded and purified as in D. Note the SDS stability of DR1 and DR4. The final yields were 12, 23 and 14%, respectively.