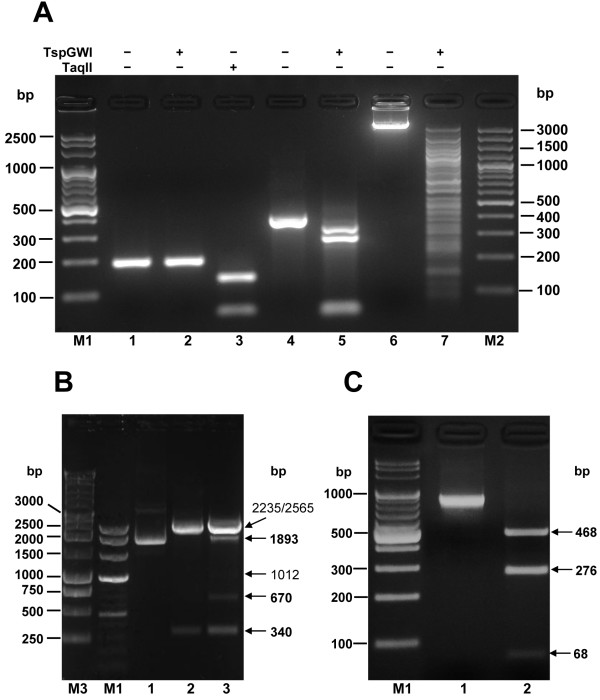
Figure 9.
TspGWI DNA substrate preferences. (A) Resistance of DNA substrate with a single target site to TspGWI cleavage. A PCR fragment of 200 bp was generated, which contains unique recognition sequences for two Thermus sp. (sub)family enzymes: TspGWI and TaqII. Lane M1, 100 bp ladder (selected bands marked); lane 1, undigested 200 bp fragment; lane 2, digestion of the 200 bp PCR fragment with TspGWI; lane 3, digestion of the 200 bp PCR fragment with TaqII; lane 4, undigested 390 bp fragment (double sites; →←); lane 5, digestion of the 390 bp PCR fragment (double sites; →←) with TspGWI; lane 6, undigested bacteriophage lambda DNA; lane 7, digestion of bacteriophage lambda DNA with TspGWI; lane M2, 100 bp DNA ladder (selected bands marked). (B) Restoration of cleavability of the 200 bp PCR fragment by cloning into pUC19. The PCR fragment refractory to TspGWI cleavage was cloned into pUC19 (yielding pUC19-200), containing two existing TspGWI sites. The complete digestion pattern includes 340, 670 and 1893 bp restriction fragments (marked in bold). The partial digestion pattern (all 3 sites) includes 340, 670, 1010, 1893, 2232 and 2562 bp. The fragments obtained are marked with horizontal arrows. Lane M3, 1 kb ladder (selected bands marked); lane M2, 100 bp ladder; lane 1, control undigested pUC19 DNA; lane 2, pUC19 DNA digested with 3 u TspGWI; lane 3, pUC19-200 digested with 3 u TspGWI. (C) Susceptibility of DNA substrate with double divergent sites to TspGWI cleavage. A PCR fragment of 816 bp was generated, which contains two divergent recognition sequences TspGWI, separated by 40 bp spacer DNA. Lane M, 100 bp DNA ladder (selected bands marked); lane 1, undigested 816 bp PCR substrate; lane 2, digestion of the 816 bp DNA with 3 u TspGWI.