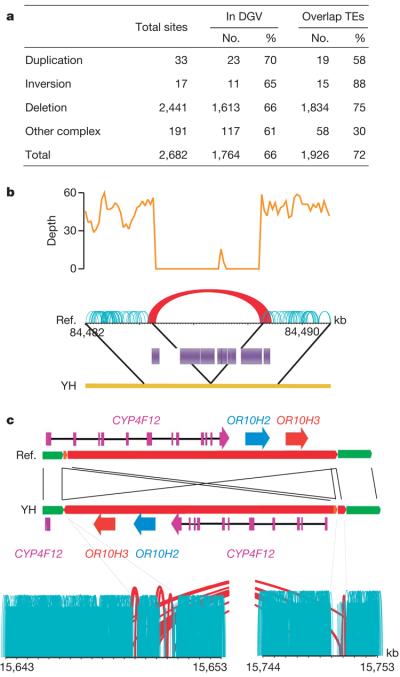
Figure 3. Summary of structural variations.
a, Abundance of each class of structural variation. The overlap with known structural variations in the DGV (http://projects.tcag.ca/variation/) and with transposons (transposable elements, TEs) was calculated. About 34% of our identified structural variations are novel (having less than 10% of a portion of the YH structural variations overlapping with structural variations in the DGV). Transposable elements are a major component of the identified deletions, with Alus and LINEs involved in 49% and 34% of the deletions, respectively. b, An example of a deletion of a transposon complex on YH chromosome 1. The sequencing depth by both single-end and paired-end reads are shown. Normally aligned paired-end reads are shown in green, whereas abnormally aligned paired-end reads, which have unexpected long insert sizes or an incorrect orientation relationship, are shown in red. c, An example of an inversion on YH chromosome 19. Local assembly showed that a 102,405-bp fragment was inverted and reinserted in the genome. There are three genes in this sequence fragment, and the last exon of gene CYP4F12 was destroyed by this inversion event.