Figure 4.
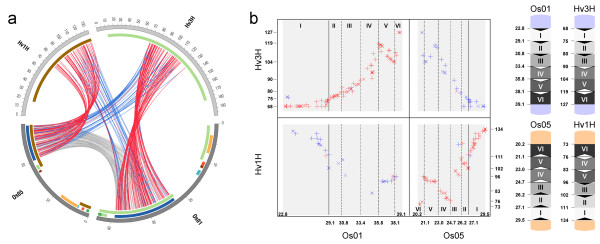
Close up view on one ancestral duplicated block (AD_1) involving Hv1H/Hv3H and Os05/Os01. a) A large degree of synteny existed between barley chromosome Hv1H and rice chromosome Os05 as well as between Hv3H and Os01 (indicated by brown and green inner bars). Grey-colored lines are shown between 354 segmentally duplicated rice genes located in the largest out of 7 duplicated segments between Os01 and Os05 (represented by colored bars in the outside lane). Best sequence homologies of barley ESTs to rice genes (putative orthologs) are represented by red and second best sequence homologies (putative paralogs) by blue connecting lines, respectively. b) Local chromosome inversions were identified in the corresponding orthologous regions of AD_1 by plotting the coordinates of mapped barley ESTs and the best (red) and second-best (blue) rice homologs against each other. Boarders of analogous subsegments I – VI corresponded to duplicated rice genes and are indicated by vertical lines. Although I – VI did not display inversions by themselves within the rice genome, inversions could be identified according to barley-rice synteny as represented schematically on the right panel of the figure. Positions are in Mb for rice and in cM for barley. Following gene pairs were used to define subsegments: LOC_Os01g50030 – LOC_Os05g47540, LOC_Os01g53079 – LOC_Os05g45320, LOC_Os01g57220 – LOC_Os05g42330, LOC_Os01g61420 – LOC_Os05g39380, LOC_Os01g65100 – LOC_Os05g35650.