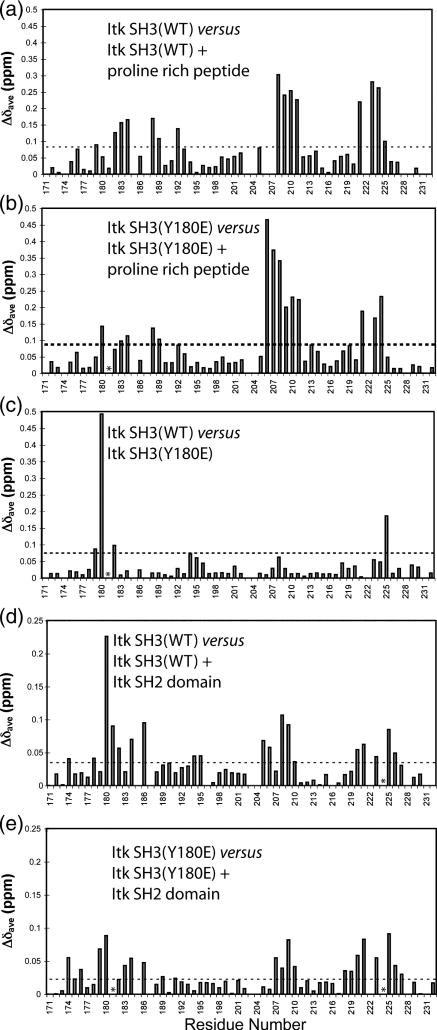
Figure 4. Chemical shift perturbations induced by Itk SH3 Y180E mutation and ligand binding.
Purified 15N labeled Itk SH3 WT, or SH3 Y180E mutant were titrated with increasing amounts of proline rich peptide or Itk SH2 domain. The average chemical shift (Δδave) for each amide is plotted against the residue number of Itk. The horizontal dashed lines represent the mean plus one standard deviation for each titration. Residues that exhibit chemical shift changes above this line are considered significant and are mapped onto the structure of the SH3 domain in Figure 5. (A) Chemical shift changes upon binding of proline rich peptide to wild type 15N Itk SH3; (B) Binding of proline rich peptide to Y180E 15N Itk SH3 mutant; (C) Chemical shift differences between wild type Itk SH3 and the Y180E Itk SH3 mutant. (D) Chemical shift changes upon binding of Itk SH2 domain to wild type 15N Itk SH3; (E) Binding of Itk SH2 domain to the Y180E 15N Itk SH3 mutant. Asterisks indicate resonances that broaden beyond detection upon ligand binding. Such residues are included in the surface maps shown in Figure 5.