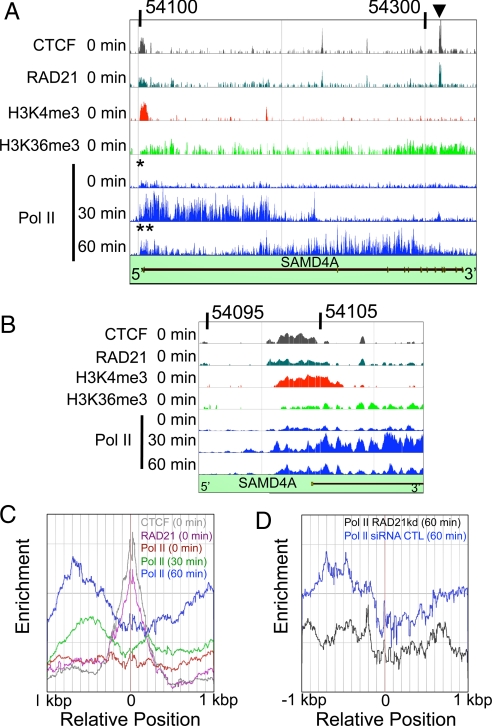
Fig. 4.
Stalling of Pol II analyzed using chromatin immunoprecipitation. HUVECs were stimulated with TNFα and harvested after 0, 30, and 60 min; then binding of CTCF, RAD21, modified histones (H3K4me3, H3K36me3), and elongating Pol II (phospho-Ser-5 modification) to SAMD4A was analyzed by ChIP-chip (CTCF, RAD21, H3K4me3, H3K36me3, and Pol II). Numbers on top of A and B show the location of the genomic region of Chr14 (SAMD4A) analyzed. (Vertical axes) Enrichment of binding. (A) Binding to SAMD4A. CTCF and RAD21 are often found together, consistent with the binding of a functional insulator complex. Asterisk shows where (engaged) Pol II binds at 0 min near the TSS, suggesting that it might be paused or poised. Double asterisk shows where Pol II binds near the TSS at 60 min. Arrowhead shows representative colocalization site of RAD21 and CTCF ≈210 kilonucleotides downstream of the TSS. (B) Binding at the TSS of SAMD4A. H3K4me3 and CTCF/RAD21 bind in/around the TSS. At 0 min, engaged Pol II also binds to this region. At 30 min, Pol II binding spreads into the gene, and after 60 min it becomes more concentrated around the TSS again. (C) Enrichments (number densities) of engaged Pol II near 35 sites distant from the TSS that were marked by bound RAD21 (pink) and CTCF (gray). At 0 min (red line), Pol II binds symmetrically around the RAD21/CTCF. At 30 min (green line), the amount of Pol II binding increases significantly. At 60 min (blue line), Pol II binding increases further and becomes concentrated upstream of the RAD21/CTCF; this is consistent with polymerase stalling. (D) An 80% reduction in RAD21 levels (achieved using siRNA) at 60 min (black) destroys the accumulation 5′ of the RAD21/CTCF site that is seen with a control siRNA (blue).