Figure 2.
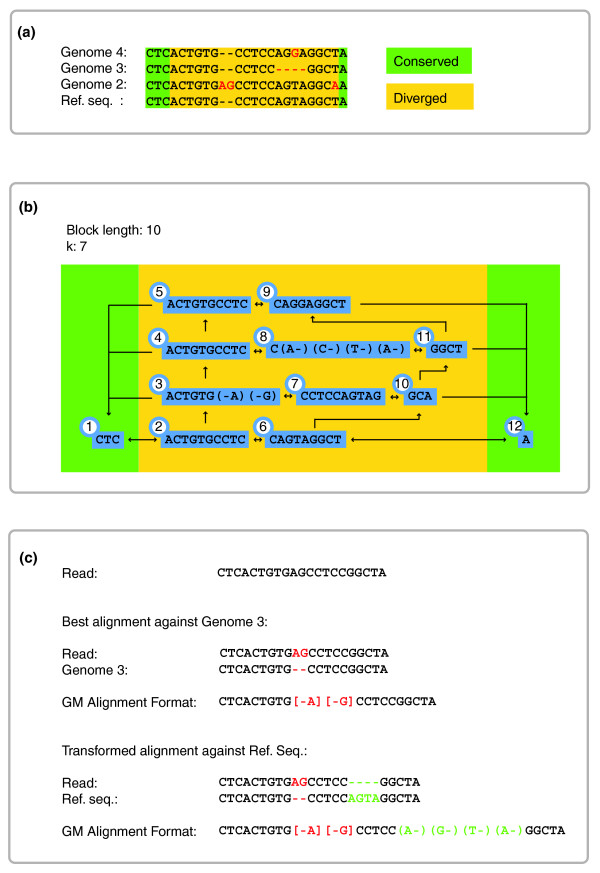
GenomeMapper's graph index structure. (a) Examples of orthologous sequences in four divergent genomes. Sequences at the beginning and end of each fragment are shared (underlaid with green boxes). Divergent regions start k-1 positions (in this case, six positions) before the first true variable position, to account for the k-mer length used for the hash-key calculation. (b) Graph structure created by these sequences, with k-mer length 7, and maximal block length of 10 (instead of 256) for reasons of illustration. The number attached to each block is its unique identifier. Note that blocks do not occupy their maximal block length after an indel, exemplified by blocks 3 and 8. Blocks 1 and 12 correspond to sequences identical in all four genomes and are present only once in the index structure. Arrows between the blocks visualize the edges between the nodes in the genome graph as they are stored in the block table [see Table S1 in Additional data file 1]. (c) Alignment of a read against the most similar genome, Genome 3, with a 2-bp insertion. Although the insertion also is observed in Genome 2, the 4-bp deletion downstream in Genome 3 makes the read more similar to it than to Genome 2. The transformed alignment of the read against the original reference sequence (Ref. seq.) includes the 4-bp deletion (as supported by Genome 3) given in parentheses (green), whereas the 2-bp insertion (which is supported neither by Genome 3 nor by the reference sequence) is annotated like a mismatch by using square brackets.