Table 1.
Performance of HM-SVM versus other methods on all data sets
| Data set | Method | Specificity+ (random)a | Sensitivity+ (random)b | F1 | Accuracy | MCC | AUC | Time (s)c |
|---|---|---|---|---|---|---|---|---|
| Hetero-complex Id | ANN | 37.6% (28.1%) | 59.4% (16.7%) | 46.0% | 60.9% | 18.9% | 64.5% | 326 |
| SVM | 38.4% (28.1%) | 59.8% (16.8%) | 46.8% | 61.8% | 20.2% | 65.4% | 179461 | |
| CRF | 42.6% (28.1%) | 55.2% (15.5%) | 48.0% | 66.5% | 24.4% | 65.3% | 12151 | |
| HM-SVM | 44.9% (28.1%) | 56.0% (15.7%) | 49.8% | 68.3% | 27.4% | 69.5% | 356 | |
| Homo-complex I | ANN | 39.0% (27.0%) | 58.4% (15.8%) | 46.6% | 63.9% | 22.1% | 67.0% | 586 |
| SVM | 39.6% (27.0%) | 61.9% (16.7%) | 48.3% | 64.2% | 24.2% | 68.6% | 224979 | |
| CRF | 45.1% (27.0%) | 59.2% (16.0%) | 51.2% | 69.5% | 30.2% | 67.6% | 16961 | |
| HM-SVM | 45.4% (27.0%) | 60.0% (16.2%) | 51.7% | 69.7% | 30.9% | 72.2% | 588 | |
| MixeI | ANN | 40.3% (27.5%) | 51.4% (14.1%) | 44.7% | 65.4% | 20.8% | 65.8% | 1242 |
| SVM | 39.5% (27.5%) | 61.5% (16.9%) | 48.1% | 63.6% | 23.3% | 67.6% | 831579 | |
| CRF | 44.3% (27.5%) | 57.5% (15.8%) | 49.9% | 68.4% | 28.0% | 66.8% | 28364 | |
| HM-SVM | 45.5% (27.5%) | 58.0% (15.9%) | 51.0% | 69.4% | 29.7% | 71.2% | 891 | |
| Hetero-complex IIf | ANN | 45.9% (34.9%) | 60.5% (21.1%) | 52.1% | 61.3% | 21.3% | 65.8% | 604 |
| SVM | 47.9% (34.9%) | 61.6% (21.5%) | 53.9% | 63.2% | 24.6% | 67.7% | 160625 | |
| CRF | 51.6% (34.9%) | 57.6% (20.1%) | 54.3% | 66.3% | 28.0% | 67.3% | 13441 | |
| HM-SVM | 54.0% (34.9%) | 56.7% (19.8%) | 55.3% | 68.0% | 30.5% | 70.7% | 464 | |
| Homo-complex II | ANN | 43.9% (32.3%) | 66.7% (21.5%) | 52.8% | 61.5% | 24.1% | 68.1 | 856 |
| SVM | 47.1% (32.3%) | 63.1% (20.4%) | 54.0% | 65.2% | 27.7% | 70.2% | 554054 | |
| CRF | 52.5% (32.3%) | 59.7% (19.3%) | 55.9% | 69.5% | 32.9% | 68.7% | 18124 | |
| HM-SVM | 53.3% (32.3%) | 60.1% (19.4%) | 56.5% | 70.1% | 34.0% | 73.4% | 851 | |
| Mix II | ANN | 46.5% (33.3%) | 53.4% (17.9%) | 49.4% | 63.7% | 21.7% | 65.8% | 1260 |
| SVM | 47.5% (33.3%) | 62.3% (20.8%) | 53.9% | 64.5% | 26.5% | 69.2% | 1316103 | |
| CRF | 52.2% (33.3%) | 58.6% (19.5%) | 55.2% | 68.3% | 30.9% | 68.1% | 856765 | |
| HM-SVM | 53.6% (33.3%) | 58.6% (19.6%) | 56.0% | 69.3% | 32.6% | 72.4% | 1320 | |
Specificity+ = TP/(TP+FP); Sensitivity+ = TP/(TP+FN); F1 = 2 × Specificity+ × Sensitivity+/(Specificity++Sensitivity+); Accuracy = (TP+TN)/(TP+TN+FP+FN); MCC = (TP × TN-FP × FN)/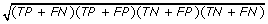 ; AUC: Area Under ROC Curve [61]. Where TP is the number of true positives (residues predicted to be interface residues that actually are interface residues); FP the number of false positives (residues predicted to be interface residues that are in fact not interface residues); TN the number of true negatives; FN the number of false negatives.
; AUC: Area Under ROC Curve [61]. Where TP is the number of true positives (residues predicted to be interface residues that actually are interface residues); FP the number of false positives (residues predicted to be interface residues that are in fact not interface residues); TN the number of true negatives; FN the number of false negatives.
aValues in parentheses are randomly predicted values. The specificity+ of random prediction is calculated as: the total number of interaction sites residues/the total number of residues.
bValues in parentheses are randomly predicted values. The sensitivity+ of random prediction is calculated as: the total number of predicted residues as interaction sites by each method/the total number of residues.
cThe total running time (second) for 5-fold cross-validation, including training and testing.
dType I data set with minor interface as negative samples.
eThe mixed data set of hetero-complexes and homo-complexes.
fType II data set with minor interface as positive samples.
