Figure 4. Comparison of the SXT/R391 core genome with the genome of pIP1202 and defining the minimal functional SXT/R391 gene set.
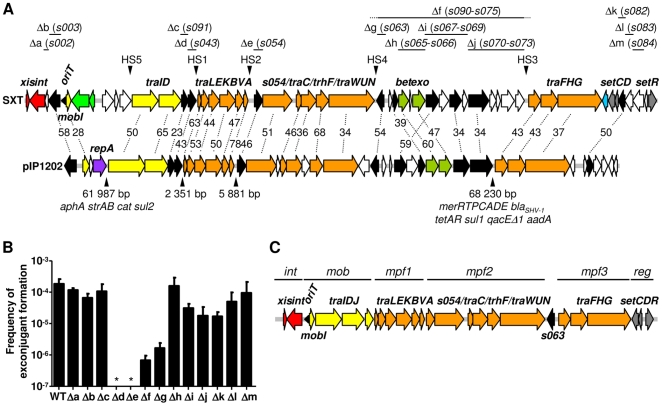
(A) Alignment of the conserved core genes of SXT/R391 ICEs with the genome of the IncA/C conjugative plasmid pIP1202 from Yersinia pestis. The top line shows the same core ICE genes shown in Figure 2A. ORFs are color coded as follows: DNA processing, yellow; mating pair formation, orange; DNA recombination and repair, green; integration/excision, red; replication, purple; regulation, gray; entry exclusion, blue; homologous genes of unknown function, black; genes without corresponding counterparts in ICEs and pIP1202, white. Numbers shown in the middle represent % identity between the orthologous proteins encoded by SXT and pIP1202 [GenBank:NC_009141]. The positions of the hotspots in SXT/R391 ICEs are marked by downward pointing arrowheads. For pIP1202, the size of the sequences (which include IncA/C backbone DNA as well as variable DNA) found at these locations as well as resistance markers are indicated by upward pointing arrowheads. aphA, aadA and strAB confer resistance to aminoglycosides. sul1 and sul2 confer resistance to sulfonamides. cat, blaSHV-1, tetAR, qacED1 and merRTPCADE confer resistance to chloramphenicol, β-lactams, tetracyclines, quaternary ammonium compounds and mercury ions, respectively. Detailed descriptions of the conserved backbone of the IncA/C conjugative plasmids have been published elsewhere [48],[50]. Regions that were deleted from SXT to investigate the function of genes of unknown function (see panel B) are indicated with straight lines. Dotted lines indicate that the deletion included DNA in the adjacent hotspots. (B) Influence of deletion of genes of unknown function on the frequency of SXT transfer. The mean values and standard deviations from three independent experiments are shown. * indicates that the frequency of transfer was below the detection level (<10−8). Deletion mutants SXTΔa, SXTΔk and SXTΔl, transferred at frequencies that were not significantly different from that of wild-type SXT (data not shown). (C) Proposed minimal set of genes necessary for a functional SXT/R391 ICE. int, integration/excision module; mob, DNA processing module; mpf, mating pair formation modules; reg, regulation module.