FIG. 2.
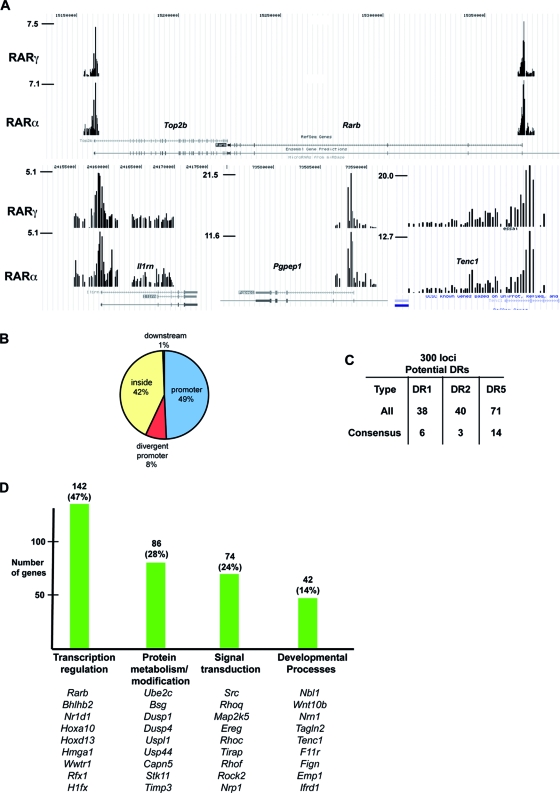
Identification of RAR binding sites in MEFs. (A) Graphic representation of tandem Flag ChIP-chip results on cells expressing tagged RARα and RARγ in the UCSC web browser (http://genome.ucsc.edu/) at the indicated loci. The values on the y axis show the normalized immunoprecipitate/input ratio. (B) Pie chart representation of the locations of RAR binding sites relative to the TSS. (C) Summary of the presence of classical DR-type RAR binding motifs at ChIPed loci. The total number of potential sites with one mismatch is shown along with the number of consensus sites. (D) Summary of gene ontology analysis of target genes. The number of genes in each category is shown along with representative examples.