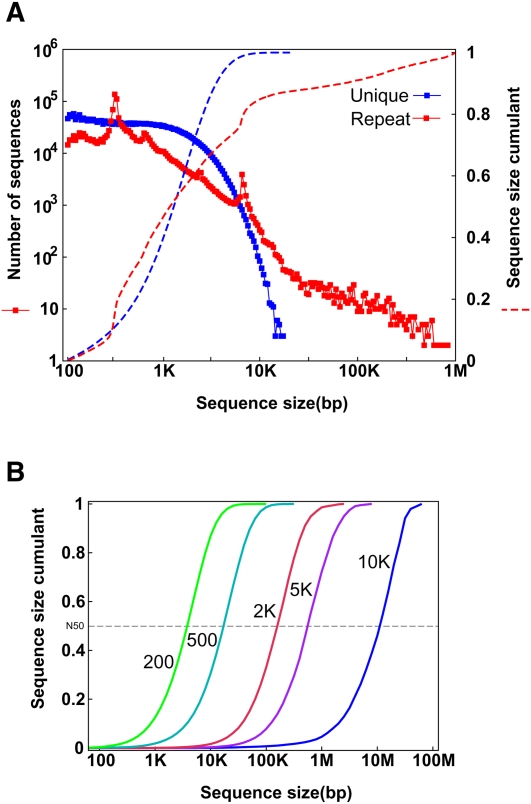
Figure 1.
(A) Length distribution of unique and repeat sequence clusters in the human genome. At each chromosomal location, we checked the frequency of the 25-mer in the whole human genome. If it appeared once, we defined it as unique; otherwise it was considered a repeat 25-mer. The regions were then merged as unique clusters and repeat clusters, and those small unique clusters (<100 bp) inside repeat clusters were defined as repeats. (B) Sequence length distribution of an ideal assembly with each insert-sized paired-ends. The repeat clusters with lengths smaller than the assumed insert size of paired-ends were crossed and the unique clusters were merged. These unique clusters represent the ideal assembly using the paired-ends.