Table 3.
EIN with experimental NOE data set from BMRB
| Case | Trial # Table S3 | N peaks | Runtime | T NOE | Additional data |
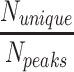 (%) (%) |
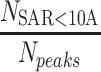 (%) (%) |
Accuracy (%) |
|---|---|---|---|---|---|---|---|---|
| 1 | – | 243 | 18 h | 2 | 28 | 53 | 99.2 | |
| 2 | 4 | 243 | 107 h | 3 | CS | 19 | 34 | 100 |
| 3 | 5 | 243 | 6 h | 2 | CS | 30 | 53 | 99.2 |
| 4 | 6 | 243 | 6 days | 2 | CS | 31 | 70 | 99.2 |
| 5 | 26 | 243 | 50 min | 3 | CS + RDC | 63 | 84 | 100 |
| 6 | 30 | 243 | 5 min | 3 | CS + CScarbon | 42 | 80 | 100 |
| 7 | 31 | 243 | 1 s | 3 | –“–, refined | 66 | 83 | 99.6 |
| 8 | 33 | 243 | 5 min | 3 | CS + CScarbon + RDC | 67 | 84 | 100 |
| 9 | 34 | 243 | 1 s | 3 | –”–, refined | 80 | 85 | 99.2 |
| 10 | – | 253 | – | – | CA(i → i − 1) with MARS | 7 | – | 100 |
| 11 | – | 253 | – | – | NOE + CS with NOEnet and CA(i → i − 1) with MARS | 97.2 | – | 99.2 |
N
NOEs = 407 experimental unambiguous NOEs have been used. The X-ray structure 1ZYM was used, yielding N
dist = 1034 distances below 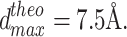 NOE classes and NOE outliers have been used. Two sets of RDCs were simulated from 1ZYM as described in material and methods. T
NOE: maximum number of allowed NOE outliers for an arbitrary matching. Case 1 was reported in (Stratmann et al. 2009)
NOE classes and NOE outliers have been used. Two sets of RDCs were simulated from 1ZYM as described in material and methods. T
NOE: maximum number of allowed NOE outliers for an arbitrary matching. Case 1 was reported in (Stratmann et al. 2009)
