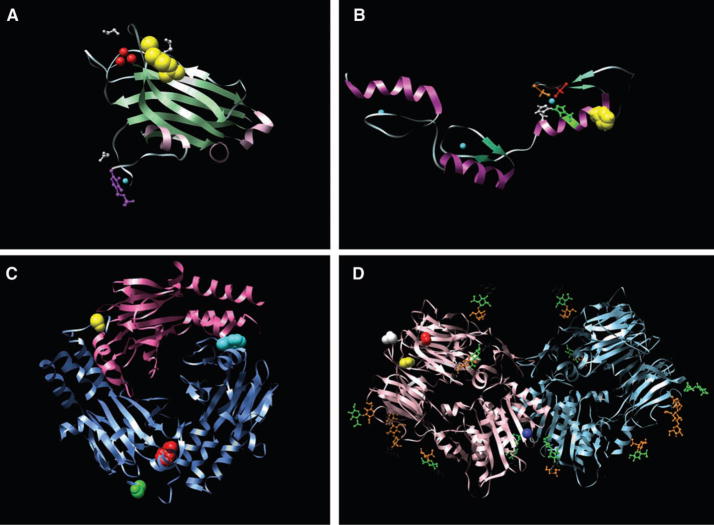
Fig. 1.
Examples of structural models of mutations. (A). The x-ray crystal structure of the C2 domain of protein kinase C γ (PKCG) [Protein Data Bank identification number (PDBID) 2UZP]. R252 (41) is shown as yellow space-fills; Ca2+ ions are shown as red spheres. The ligands 1,2-ethanediol and pyridoxal-5-phosphate are shown in white and purple ball-and-stick representations, respectively. The R252→H252 (R252H) mutation could reduce the membrane binding of the C2 domain of PRKCG and thereby affect function. (B) The nuclear magnetic resonance solution structure of the three tandem repeats of zf-C2H2 domains from human Kruppel-like factor 5 (KLF5) (PDBID 2EBT). H389 is shown as yellow space-fills; Zn2+ ions are shown as cyan spheres. The residues comprising the C2H2 group that coordinate the nearby Zn2+ ion are shown as ball-and-stick representations, H393 and H397 are shown in green and white, whereas C380 and C375 are shown in orange and red. The mutation at position 389 (H389N) may disrupt the structure of the zinc finger or nearby zinc coordination site. (C) The x-ray crystal structure of the heterotrimer of SMAD3 (two subunits shown as blue ribbons) and SMAD4 (one subunit shown as pink ribbons) (PDBID 1U7F). The residues corresponding to two of the mutant positions (F260S and S422F, shown as red and yellow space-fills, respectively, in chain A) are located at interfaces and could perturb Smad3-Smad3 or Smad3-Smad4 interactions. In chain B, F260 is shown as cyan space-fills and S422 as green space-fills. (D) The x-ray crystal structure of the extracellular domain of human DPP6 as a homodimer (PDBID 1XFD). Two of the mutated residues found in this study, T409I (shown as red space-fills) and D475N (shown in yellow space-fills) are in spatial proximity and are close to one of the glycosylation sites, N471 (shown as white space-fills). These mutations fall in the β-propeller domain of the protein (residues 142 to 322 and 351 to 581) thought to be involved in protein-protein interactions. The A778T mutation (shown as blue space-fills) falls in the */β hydrolase domain (residues 127 to 142 and 581 to 849) and is close to the homodimer region of the protein and could perturb the homodimer association. Carbohydrates with glycosylation sites are shown in stick representation. Images created with UCSF Chimera version 1.2422 for Linux (42).