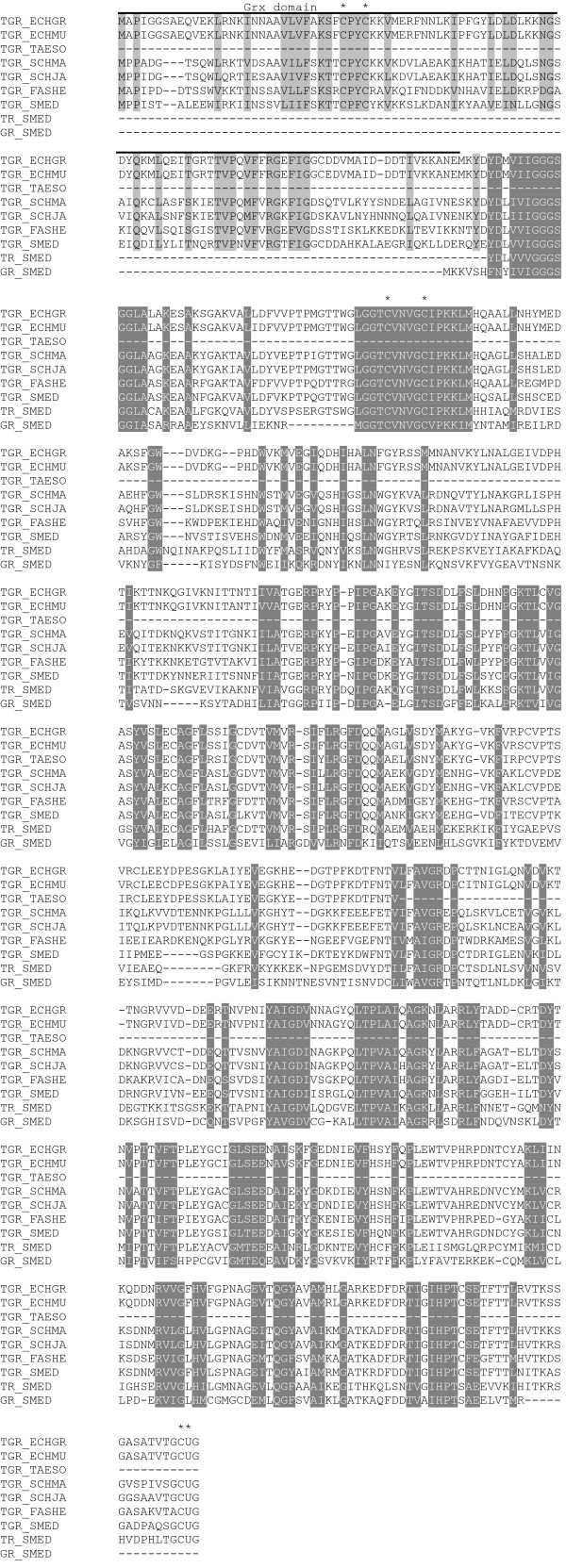
Figure 1.
Amino acid sequence alignment of TR, GR and TGRs of platyhelminths. Sec is indicated by U. The position of the redox active residues in the sequences is indicated by a star. Conserved residues in all proteins are highlighted in dark grey, conserved residues in the Grx domain of TGRs are highlighted in light grey. Location of the Grx domain is indicated above the sequence. ORFs for TGR, TR and GR from S. mediterranea (SCHME) genome (assembly 31) were predicted in the contigs 000676, 000203 and 001663, respectively. ORF for E. multilocularis (ECHMU) TGR was assembled from contigs 0007357 and 0007358. Full-length TGR sequences of S. mansoni (SCHMA),S. japonicum (SCHJA), E. granulosus (ECHGR), and F. hepatica (FASHE) were retrieved from Genebank (gb|AAK85233.1|AF395822_1, gb|AAW25951.1, emb|CAM96615.1, and gb|AAN63052.1, respectively). T. solium (TAESO) TGR partial sequences were retrieved from the EST repository at Genebank (gb|EL757065.1 and gb|EL743442.1). The putative mitochondrial leader peptide of S. mediterranea TGR is not included in the sequence, neither leader peptide variants of Schistosoma and Echinococcus TGRs. Sequences were aligned with Clustal W2 [29], with final manual adjustment after inspection.