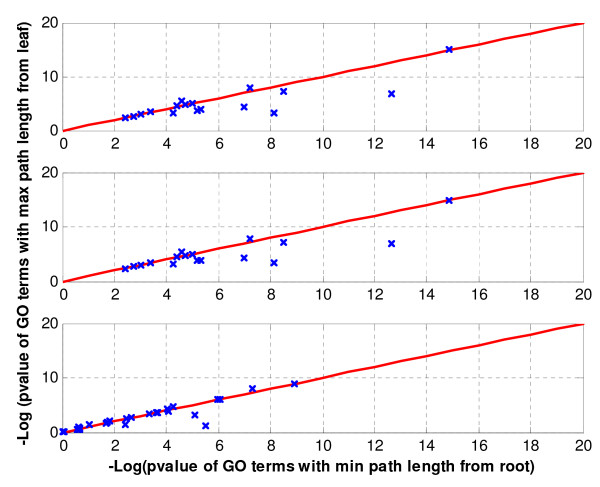
Figure 6.
Comparative analysis of the max path length from leaf (Strategy I) and the min path length from root (Strategy II) approaches in terms of p-value of each GO term. Each row represents one of the three datasets: Saccharomyces cerevisiae amino acid starvation data, Saccharomyces cerevisiae cell cycle data, and Brassica napus data respectively. The y-axis represents the minus log P-value for GO enrichment using the Strategy I (0, the most specific term by itself; no parent is considered). The x-axis represents the minus log P-value for GO enrichment using Strategy II (3, a default in several tools). Points above the central diagonal line represent the most significant GO-terms in Strategy I, whereas points below the line represent categories with most significant terms in Strategy II. As can be seen, Strategy II has slightly larger number of significant GO terms.