Figure 3.
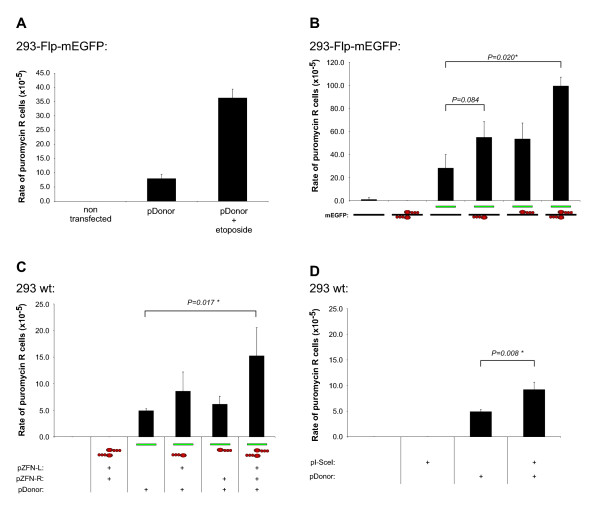
Analysis of illegitimate genomic integration of the donor plasmid in ZFN treated cells. (A) Effect of etoposide treatment on the levels of illegitimate integration of the PuroR expression cassette from the pDonor in the genome of 293-Flp-mEGFP cells. After transfection with pDonor, cells were either left untreated or treated with etoposide before selection with puromycin in order to quantify the levels of illegitimate integration. As a control for the selection conditions, non-transfected cells were also included. Each bar represents the mean rate of puromycin resistant (R) cells ± SD from three parallels. (B) Quantification of illegitimate integration in 293-Flp-mEGFP cells after transfection with pDonor along with pZFN-L and pZFN-R. Controls included non-transfected cells and cells transfected with pZFNs or pDonor individually as indicated below the bars. The rates of illegitimate integration were normalized according to variations in PE. Statistical P values (paired t-test) comparing the means of the indicated bars are shown and an asterisk marks significant difference (P < 0.05). (C) Analysis of illegitimate genomic integration of the pDonor in wt 293 cells that did not enclose the mEGFP target sequence in their genome. Cells were transfected with pDonor, pZFN-L and pZFN-R as indicated below the bars and analyzed as in B. (D) Analysis of illegitimate genomic integration of the pDonor in wt 293 cells after transfection with an I-SceI endonuclease expression construct (pI-SceI). Cells were transfected with pDonor together with 100 ng of pI-SceI. The frequency of illegitimate integration was quantified as in B.