Figure 2.
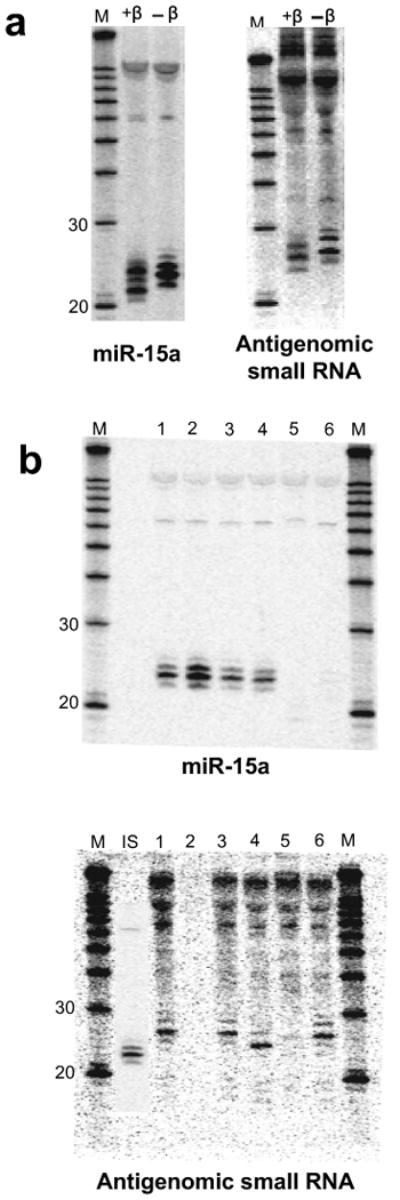
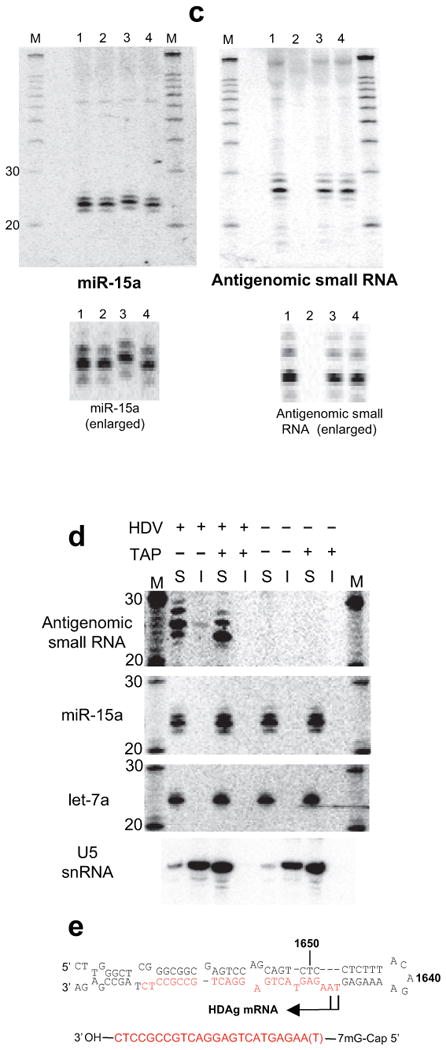
The antigenomic HDV small RNA is 2′-3′ hydroxylated and has an mRNA-like 5′ cap (Northern Blot, 293 cells, RNA induction). (a) 3′ end by β-elimination. The mobility of the HDV small RNA is increased following β-elimination. miR-15a: 2′-3′ hydroxylated positive control; +β: +β-elimination; -β: untreated RNA. (b) 5′ end by enzymatic analysis. 1: mock-treated (+HDV); 2: mock-treated (no HDV); 3: PNK (+HDV); 4: Decapping enzyme (TAP; +HDV); 5: T4 RNA Ligase (+HDV); 6: Terminator Exonuclease (+HDV). The size of the HDV small RNA was estimated to be ∼24nt based on the largely 22nt, 5′ phosphorylated miR15-a shown in the inset (IS). (c) Confirmation that the 5′ end of the HDV small RNA is capped, not triphosphorylated (enlarged image to emphasize changes in gel mobility for miR-15a, but not HDV small RNA). 1: mock-treated (+HDV); 2: mock-treated (no HDV); 3: Antarctic Phosphatase (+HDV); 4: Antarctic Phosphatase followed by T4 PNK (+HDV). (d) RNA immunoprecipitation with anti-2,2,7-trimethylguanosine antibody K121. The immunoprecipitation efficiency of the HDV small RNA, U5 snRNA (positive control) and microRNAs miR-15a and let-7a (negative controls) was analysed by Northern blot. ‘S’: supernatant; ‘I’: IP fraction. (e) Predicted structure of the HDV small RNA. The various RNAs in a-d were detected after stripping and rehybridisation to the same blot. M: RNA marker.
