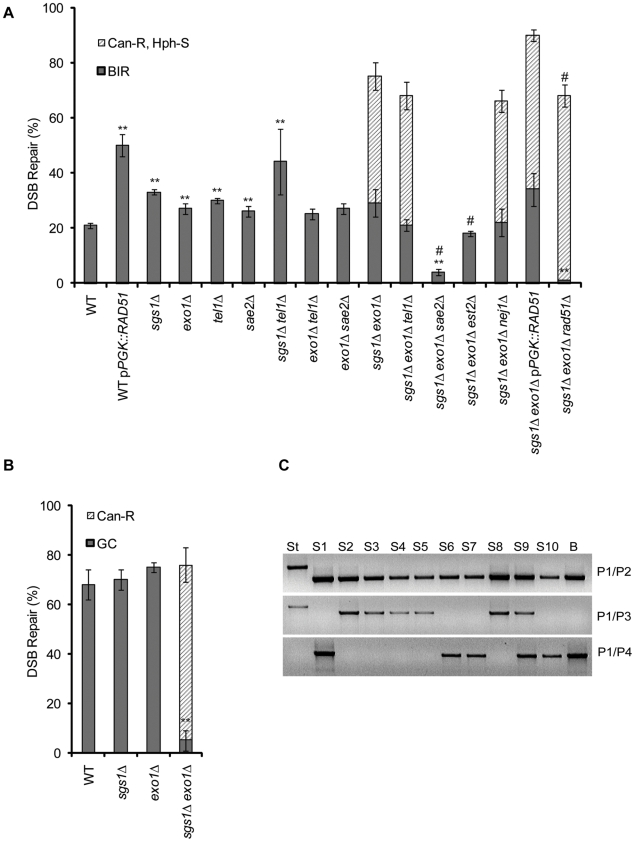
Figure 4. The effect of sgs1Δ exo1Δ on the viability and repair product in BIR and GC.
(A) The viability and phenotypic characterization of wild type (WT), sgs1Δ, exo1Δ, tel1Δ, sgs1Δsgs1Δ exo1Δ, pPGK::RAD51 and indicated double and triple mutant combination cells following a DSB in the BIR assay. BIR colonies (CanS HphS) represent those that have repaired the DSB by BIR while CanR HphS colonies represent those that have a truncated chromosome. Data are the mean ±s.e.m. Values marked with asterics or number sign are statistically significant (*represents p < 0.05, ** represents p < 0.01 compared to wild type BIR. # represents p < 0.05 to the sgs1Δ exo1Δ CanR HphS colonies). (B) The viability and phenotypic characterization of cells following a DSB in the GC assay. HR colonies (CanS) represent those that have repaired by Homologus Recombination (either BIR or GC) while CanR colonies represent those that have a truncated chromosome. Data are the mean ±s.e.m. ** represents p < 0.01 compared to wild type. (C) Repair of CanS colonies in the GC assay as monitored by PCR. Included are the starting GC strain (ST), ten CanS colonies (S1–S10) and a colony that has repaired by BIR (B). PCR with primers P1 and P2 detects the starting band and shift to smaller size upon repair into the CAN1 sequences if repair occurs either by GC or BIR. PCR of primers P1 and P3 monitors retention of the distal end of the DSB and is indicative of repair by GC. PCR with primers P1 and P4 monitors repair specific to BIR (see Figure 1).