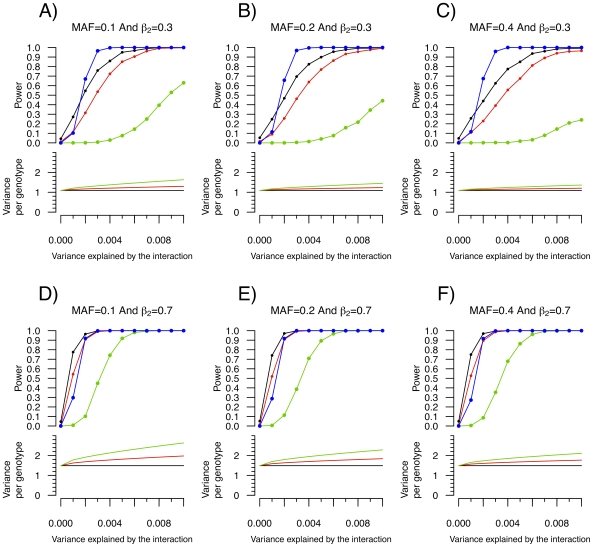
Figure 3. Power to detect a SNP–covariate interaction effect as function of the proportion of variance explained by the interaction.
Each condition was simulated 1,000 times with 15,000 individuals according to the model shown in Figure 1. MAF refers to the minor allele frequency of SNP1 and β2 refers to the β coefficient of the C1-Y association. In all cases, SNP1 had no marginal effect (i.e. β1 = 0). Upper panel: Power to identify SNP1 as an “interacting” SNP using Levene's test with a P-value threshold of 0.05 (black), 0.01 (red) and 1.5×10−7 (green; to account for 340,000 tests using Bonferroni correction). Also shown is the power to detect the interaction itself with a linear regression interaction P-value cut-off of 1.5×10−7 (blue). Lower panel: The variance per genotype is illustrated as a function of the fraction of the total variance of the quantitative trait explained by the interaction. The homozygous major allele genotype is drawn in black, the heterozygous genotype in red and the homozygous minor allele genotype in green.