Figure 4.
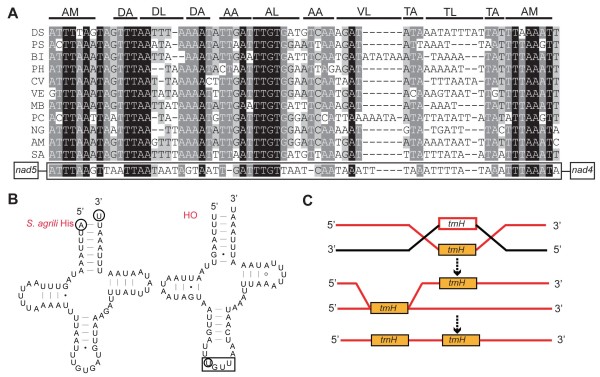
Mechanism of trnH remote inversion in Spathius agrili mitochondrial genome. (A) Presumed pseudo-trnH sequence (HO) and 10 hymenopteran trnH sequences are aligned according to their secondary structures. AM: Accepter arm, DA: D-loop arm, DL: D-loop, AA: Anticodon arm, AL: Anticodon loop, VL: Variable loop, TA: T Ψ C arm, TL: T Ψ C loop. (B) Secondary structure trnH is predicted in tRNAscan-SE search server [82] and HO is predicted manually. The inserted uracil in the anticodon is showed by a circle. (C) Recombination of two strands with opposite orientations leads the inversion of trnH, and the following recombination events lead the duplication of trnH. DS: Diadegma semiclausum, PS: Primeuchroeus spp., BI: Bombus ignites; PH: Polistes humilis, CV: Cotesia vestalis, VE: Vanhornia eucnemidarum, MB: Melipona bicolor, PC: Perga condei, NG: Nasonia giraulti, AM: Apis mellifera, SA: Spathius agrili.