Abstract
Background
Identification of genes involved in adaptation and speciation by targeting specific genes of interest has become a plausible strategy also for non-model organisms. We investigated the potential utility of available sequenced fish genomes to develop microsatellite (cf. simple sequence repeat, SSR) markers for functionally important genes in nine-spined sticklebacks (Pungitius pungitius), as well as cross-species transferability of SSR primers from three-spined (Gasterosteus aculeatus) to nine-spined sticklebacks. In addition, we examined the patterns and degree of SSR conservation between these species using their aligned sequences.
Results
Cross-species amplification success was lower for SSR markers located in or around functionally important genes (27 out of 158) than for those randomly derived from genomic (35 out of 101) and cDNA (35 out of 87) libraries. Polymorphism was observed at a large proportion (65%) of the cross-amplified loci independently of SSR type. To develop SSR markers for functionally important genes in nine-spined sticklebacks, SSR locations were surveyed in or around 67 target genes based on the three-spined stickleback genome and these regions were sequenced with primers designed from conserved sequences in sequenced fish genomes. Out of the 81 SSRs identified in the sequenced regions (44,084 bp), 57 exhibited the same motifs at the same locations as in the three-spined stickleback. Di- and trinucleotide SSRs appeared to be highly conserved whereas mononucleotide SSRs were less so. Species-specific primers were designed to amplify 58 SSRs using the sequences of nine-spined sticklebacks.
Conclusions
Our results demonstrated that a large proportion of SSRs are conserved in the species that have diverged more than 10 million years ago. Therefore, the three-spined stickleback genome can be used to predict SSR locations in the nine-spined stickleback genome. While cross-species utility of SSR primers is limited due to low amplification success, SSR markers can be developed for target genes and genomic regions using our approach, which should be also applicable to other non-model organisms. The SSR markers developed in this study should be useful for identification of genes responsible for phenotypic variation and adaptive divergence of nine-spined stickleback populations, as well as for constructing comparative gene maps of nine-spined and three-spined sticklebacks.
Background
Recent advances in our understanding of the physiological and molecular functions of genes have paved the road for investigating functional genomic variation associated with adaptation and speciation in the wild [1,2]. Consequently, targeting specific genes and genomic regions of interest - rather than random genomic regions - holds a great promise as a shortcut to identify genes involved in phenotypic variation and adaptive divergence [3-5]. Despite a steadily increasing number of completed genome sequences, genomic resources and tools are still very limited for the vast majority of non-model organisms. Therefore, ability to develop molecular markers in or around target genes is essential for application of this approach for non-model organisms. In addition, molecular markers associated with functionally important genes are useful in construction of comparative genetic maps, in which they can be exploited as comparative anchor tagged sequence loci [6,7].
Microsatellites or simple sequence repeats (SSRs) are highly abundant in eukaryotic genomes, accounting for 3-5% of the mammalian genomes [8,9]. Owing to their wide genomic distribution, codominant inheritance and hypervariability, they are widely recognized as one of the most powerful molecular markers in the field of genetics. As a result of the widespread use of SSRs, substantial efforts have been made to devise procedures for developing SSR markers [10,11]. In addition, cross-species transfer of SSR primers is commonly attempted in many taxa [12]. However, SSR markers developed with conventional approaches are derived from the genome more or less in a random manner. Expressed sequence tags (ESTs) are commonly used as an alternative to genomic libraries as a source of SSR markers [11]. SSR markers derived from ESTs have some advantages over those developed from genomic libraries because EST-derived markers can associate with genes of known or putative function, and they exhibit relatively high transferability between closely related species [13-17]. However, SSRs are generally much less abundant in transcribed regions than in non-transcribed regions [18-21] and found typically only in a few percentage of ESTs [15,22-26]. Besides, designing primers requires sufficient flanking sequences, resulting in a considerable reduction in number of ESTs available to develop SSR markers [23,24,26,27]. Therefore, even if a large EST database is available for a target species, ESTs have limitations as a material for development of SSR markers for specific genes.
One way to obtain SSR markers for specific genes and genomic regions in a given species is to use SSR primers developed for the closest relative with a sequenced genome. However, an obvious limitation of this approach is that mutations in SSR flanking sequences will inhibit cross-species amplification success - a problem that is likely to attenuate with an increasing divergence time [28]. In general, success of cross-species transfer is a negative function of the evolutionary distance separating the source and focal species [28-31]. Another crucial issue is related to evolution and persistence of SSRs among different species. Investigations of SSR conservation have demonstrated that several SSRs are retained not only in closely related species, but also in species that have diverged more than 100 million years ago [32-35]. Nevertheless, comprehensive surveys of SSR conservation using aligned sequences of different species have rarely been reported [36,37], making it difficult to estimate the patterns and degree of SSR conservation in different taxa.
For the reasons elaborated above, development of SSR markers for target genes and genomic regions in non-model organisms is challenging. Yet, while the closest relative with a sequenced genome is too distantly related to the focal species, one can take an advantage of the increasing number of completed genomes for different species. For instance, as often used in species for which no direct species-specific sequence information is available, conserved sequences in specific genes and genomic regions of interest can be used to design primer sequences applicable to a wide variety of organisms [e.g. [38]].
Teleosts consist of approximately 28,000 species [39], which correspond to more than half of all living vertebrates. Despite a number of features of evolutionary interest and economical importance, genomic resources and tools are still lacking for most teleost taxa. Currently, genome sequences are available for five species - zebrafish (Danio rerio), three-spined stickleback (Gasterosteus aculeatus), medaka (Oryzias latipes), spotted green pufferfish (Tetraodon nigroviridis) and fugu (Takifugu rubripes) [40]. The development of genome sequences for three-spined sticklebacks has made great contribution to an understanding of the genetic architecture of several phenotypic traits [41-44]. Because three-spined and nine-spined (Pungitius pungitius) sticklebacks exhibit similar ecological and morphological characteristics [45], these species provide an opportunity to study whether the same genes or genomic regions are responsible for phenotypic variation of certain traits and adaptive divergence in different lineages. This would facilitate a molecular understanding of the parallel evolution of these species, which have diverged more than 10 million years ago - equivalent to 5-10 millions of generations [46,47]. A potentially effective strategy to this end would be to develop SSR markers targeting functionally important genes.
The main objective of this study was to develop a large set of SSR markers targeting specific genes and genomic regions for a non-model organism - the nine-spined stickleback - in which genome sequences and ESTs are not yet available. To this end, two strategies were adopted. First, we tested cross-species utility of 158 SSR primer sets for functionally important genes originally developed in three-spined sticklebacks together with 188 SSR markers derived from genomic libraries and ESTs. Secondly, we investigated the potential utility of available sequenced fish genomes to develop SSR markers for functionally important genes in nine-spined sticklebacks. To address prospects for this approach, the patterns and degree of SSR conservation were examined in three-spined and nine-spined sticklebacks using their aligned sequences.
Results and discussion
Cross-species utility of three-spined stickleback primers
Out of the 158 SSR markers for functionally important genes (gene-based SSRs), 27 showed robust and specific amplification within the expected size range in nine-spined sticklebacks (Table 1, see also Additional files 1 and 2), resulting in a low level (17.1%) of cross-species amplification. In contrast, amplification success was 34.7% (35 out of 101) and 40.2% (35 out of 87) in the SSR markers derived from genomic libraries (genomic SSRs) and ESTs (EST-derived SSRs), respectively (Table 1). The tendency for higher amplification success with the EST-derived SSRs than with the genomic SSRs is in agreement with the results of previous studies [48-51] - finding which has been explained by high sequence conservation in coding regions [13-15].
Table 1.
Cross-species amplification of three-spined stickleback SSR primers in nine-spined sticklebacks.
| SSR type | Primer site | N | Amplified | Polymorphic |
|---|---|---|---|---|
| Gene-based | All | 158 | 27 (17.1%) | 16 (10.1%) |
| Exonic pair | 9 | 7 (77.8%) | 4 (44.4%) | |
| Intronic pair | 53 | 2 (3.8%) | 1 (1.9%) | |
| Intergenic pair | 55 | 9 (16.4%) | 7 (12.7%) | |
| Other combinations | 41 | 9 (22.0%) | 4 (9.8%) | |
| Genomic | All | 101 | 35 (34.7%) | 26 (25.7%) |
| Exonic pair | 1 | 1 (100%) | 1 (100%) | |
| Intronic pair | 34 | 14 (41.2%) | 10 (29.4%) | |
| Intergenic pair | 58 | 16 (27.6%) | 11 (19.0%) | |
| Other combinations | 6 | 3 (50.0%) | 3 (50.0%) | |
| Unknown | 2 | 1 (50.0%) | 1 (50.0%) | |
| EST-derived | All | 87 | 35 (40.2%) | 21 (24.1%) |
| Exonic pair | 14 | 7 (50.0%) | 6 (42.9%) | |
| Intronic pair | 11 | 4 (36.4%) | 1 (9.1%) | |
| Intergenic pair | 47 | 17 (36.2%) | 10 (21.3%) | |
| Other combinations | 15 | 7 (46.7%) | 4 (26.7%) |
Factors affecting cross-species amplification success were assessed using the 388 SSR markers. A hierarchical generalized linear model (GLM) revealed a significant influence of SSR type (cf. gene-based, genomic vs. EST-derived SSRs) on amplification success (F2,331 = 8.28, P = 0.016). In addition, amplification success was significantly affected by primer site (cf. exonic, intronic, intergenic vs. other combinations; F3,331 = 16.43, P < 0.001). Across the three SSR types, amplification success was high for the SSR markers in which both forward and reverse primers were located in exonic regions (62.5%, 15 out of 24), whereas it was lower if primers were located either in intronic (20.4%, 20 out of 98) or intergenic regions (26.3%, 42 out of 160; Table 1). This effect was particularly obvious for the gene-based SSRs with intronic primers, in which case the amplification success was very low (3.8%, two out of 53; Table 1). As for EST-derived SSRs, trinucleotide SSRs are the most abundant repeat motif in ESTs and tend to be found in coding regions, whereas dinucleotide SSRs are often found in untranslated regions [48]. The fact that most of the EST-derived SSRs used in our study are dinucleotide repeats (85 out of 87) suggests that a number of the EST-derived SSRs might be located in untranslated regions. While in theory EST-derived SSRs should be located in exonic regions, 54 (out of 87) SSRs were located in intergenic regions according to the Ensembl genebuild. This inconsistency could be due to artifacts such as prediction errors and contamination of cDNA libraries with genomic DNA. Nevertheless, the result that amplification success tended to be higher for the EST-derived SSRs with exonic primers (50.0%, seven out of 14) than for those with intergenic primers (36.2%, 17 out of 47) might, at least in part, result from the fact that sequence homology is less in untranslated regions and increase toward the start codon of the coding regions in related species [51].
While the effects of SSR type and primer site were significant, amplification success was not significantly associated with average primer length (F1,331 = 1.09, P = 0.297) or average GC content (F1,331 = 3.12, P = 0.077). Similarly, amplification success was independent of differences in GC content (F1,331 = 0.00, P = 0.993) and melting temperature (F1,331 = 0.53, P = 0.465) between primers within a given primer pair. In addition, there was no association between amplification success and expected PCR product size (F1,331 = 0.393, P = 0.531). While the positive effect of average melting temperature appeared to be significant (F1,331 = 5.82, P = 0.016), no clear difference of melting temperature was found among the SSR types. Based on these results, it is unlikely that differential amplification success among the three sets of SSR markers stemmed from different primer conditions. In fact, all of the SSR markers were successfully amplified under the same PCR and DNA conditions in three-spined sticklebacks. Multiple gene copies are known to exist in several functionally important genes [52-55]. Rather than primer conditions, divergence of functionally important genes might be the cause of low cross-species amplification success of the gene-based SSRs.
In the gene-based SSRs, polymorphism was found at 16 out of the 27 amplifying loci (59.3%; Table 1). This rate was similar to that observed in the genomic SSRs (74.3%, 26 out of 35) and EST-derived SSRs (60.0%, 21 out of 35; Table 1). In total, 63 out of the 97 amplified loci exhibited polymorphism in Fennoscandian populations (Additional file 1). For the amplified loci, incidence of polymorphism was independent of SSR type (GLM, F2,90 = 3.37, P = 0.185), SSR location (cf. exonic, intronic vs. intergenic regions; F2,90 = 5.11, P = 0.078) and SSR repeat motif (cf. di- vs. trinucleotide repeats; F1,90 = 0.02, P = 0.876). The relatively high proportion of polymorphic loci across the different SSR types and SSR locations suggests that several SSRs are conserved in three-spined and nine-spined sticklebacks. For the polymorphic loci of gene-based SSRs, an average of 8.8 alleles per locus (range = 2-38) were identified in the three populations (Additional file 1). This value was equivalent to that obtained in the genomic SSRs (9.3) and EST-derived SSRs (8.3; Additional file 1). Average heterozygosity varied from 0.19 to 0.55 in the gene-based SSRs, from 0.07 to 0.64 in the genomic SSRs and from 0.14 to 0.63 in the EST-derived SSRs among the three populations (Additional file 1). MICRO-CHECKER analyses did not indicate the presence of null alleles, with the possible exceptions of the CLCN7, GS1, Gac7080P and Stn18 in the Baltic Sea, the Stn127 and GAest41 in the Lake 1 and the GAest16 in the Pyöreälampi. There was no evidence for deviations from Hardy-Weinberg equilibrium at any locus in any of the populations.
In general, our results demonstrate that cross-species utility of SSR primers for functionally important genes is less efficient as compared to that of genomic and EST-derived SSR markers. This is attributed to limited amplification success rather than a low incidence of polymorphism. Therefore, the development of species-specific primers would be necessary for obtaining SSR markers for functionally important genes.
SSR conservation in sticklebacks and marker development
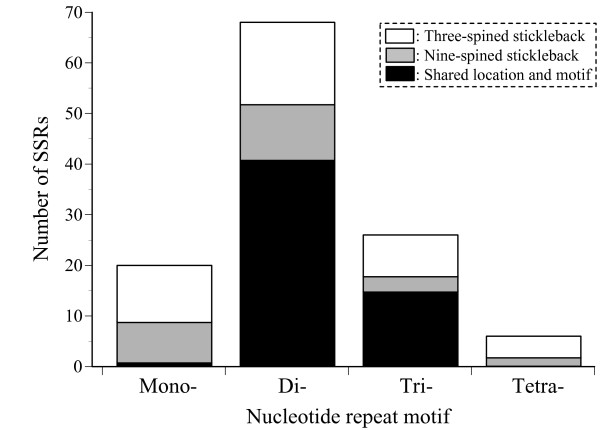
To investigate the potential utility of available sequenced fish genomes for SSR marker development in nine-spined sticklebacks, we surveyed SSRs within and around 67 functionally important genes in the three-spined stickleback genome and designed 70 primer sets for amplification and sequencing of these SSR regions using the conserved sequences determined by sequenced fish genomes (Additional file 3). The PCR product size of respective genomic regions obtained in nine-spined sticklebacks was concordant with that estimated from the three-spined stickleback genome (Additional file 4). All of the sequences of nine-spined sticklebacks for the 70 regions exhibited the highest BLAST hit scores in the target regions and high homologies to the sequences of the three-spined stickleback (Additional file 4). Out of the 70 genomic regions representing 44,084 bp, 49 contained at least one SSR in nine-spined sticklebacks (Additional file 4). The total number of SSRs observed in these regions was 81, including 9 mono-, 52 di-, 18 tri- and two tetranucleotide motifs (Figure 1). In the three-spined stickleback genome, 96 SSRs were found in the 70 homologous regions, including 12 mono-, 57 di-, 23 tri- and four tetranucleotide motifs (Figure 1). Out of the 81 SSRs found in nine-spined sticklebacks, 64 were identified at the same locations as those of the three-spined stickleback (Additional file 4). In addition, 57 out of the 64 SSRs exhibited the same motifs as those of the three-spined stickleback (Figure 1, see also Additional file 4), indicating that a large proportion of SSRs are conserved in the genomes of these species. Our results also demonstrated that SSRs with di- and trinucleotide repeat motifs are highly conserved but those with mononucleotide repeat motifs are less so (Figure 1). Hence, the level of SSR conservation may differ among SSRs differing in repeat motif type.
Figure 1.
Number of SSRs observed in 70 genomic regions in three-spined and nine-spined sticklebacks. Open, unique SSRs for three-spined stickleback; gray, unique SSRs for nine-spined stickleback; black, SSRs of shared motif and location between these species.
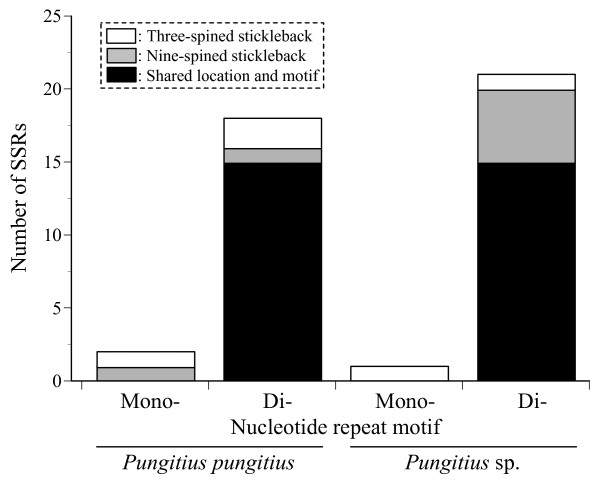
To further address SSR conservation in stickleback species, we investigated if SSRs randomly derived from genomic libraries of Pungitius species are found at the homologous genomic locations of three-spined sticklebacks. For this analysis, we used publicly available SSR and flanking sequences of Pungitius pungitius (i.e. nine-spined stickleback) [56] and Pungitius sp. [57] - so called the Omono-type, which has been regarded as an independent species from Pungitius pungitius based on the biological species concept [58]. The 13 Pungitius pungitius sequences (5,310 bp) contained one mono- and 16 dinucleotide motif SSRs. In the three-spined stickleback genome, 18 SSRs were identified in the homologous regions, including one mono- and 17 dinucleotide motifs. Out of the 18 SSRs identified in the three-spined stickleback, 15 (83.3%) exhibited the same motifs at the same locations as in Pungitius pungitius (Figure 2). In the 19 Pungitius sp. sequences (4,117 bp) containing 20 dinucleotide motif SSRs, 17 SSRs were identified in the homologous regions of the three-spined stickleback genome, including one mono- and 16 dinucleotide motifs (Figure 2). Out of the 17 SSRs identified in the three-spined stickleback, 15 (88.2%) exhibited the same motifs at the same locations as in Pungitius sp. (Figure 2). The comparative analyses of randomly selected Pungitius SSRs in the three-spined stickleback genome further indicated a high degree of SSR conservation in stickleback species.
Figure 2.
Number of SSRs observed in 32 genomic regions in three-spined sticklebacks and Pungitius species. Open, unique SSRs for three-spined stickleback; gray, unique SSRs for nine-spined stickleback; black, SSRs of shared motif and location between these species.
While several studies have reported conservation of single SSRs between different taxa [32-35,59], a comprehensive survey of SSR conservation is limited to a comparison of human (Homo sapiens) and chimpanzee (Pan troglodytes) [36,37], which have diverged six million years ago [60]. According to Vowles and Amos [37], 70% of human SSRs are homologues in chimpanzees. Our results demonstrated that a similar proportion (70% for randomly selected motifs) of SSRs are retained in three-spined sticklebacks and Pungitius species despite longer divergence time (cf. more than 10 million years) [46] and much shorter generation times (cf. one or two years) [47].
Based on the sequences obtained in nine-spined sticklebacks, species-specific primer sets were designed to amplify 58 SSRs targeting 57 functionally important genes (Table 2). Among them, polymorphism was identified at 41 loci (Table 2) in Fennoscandian populations. On average, 7.7 alleles per locus (range = 2-27) were identified across the three populations (Table 3). Average heterozygosity was 0.57 in the Baltic Sea, 0.37 in the Lake 1 and 0.06 in the Pyöreälampi (Table 3). There was no indication for the presence of null alleles, with the possible exceptions of the Ppgm40 and Ppgm50 in the Baltic Sea and the Ppgm52 and Ppgm56 in the Lake 1. Deviations from Hardy-Weinberg equilibrium were not observed at any locus in any of the populations.
Table 2.
SSR markers for functionally important genes in nine-spined sticklebacks and their polymorphism (P) in three populations.
| Gene ID | SSR marker | GenBank accession no | |||||
|---|---|---|---|---|---|---|---|
| Locus | Repeat motif | Forward primer (5'-3') | Reverse primer (5'-3') | Ta (°C) | P | ||
| ACAPRa | Ppgm1 | (CTC)12 | AGCTGCCATTTTAAATCCTCCTC | CTCACCATGATGGAAGCCAC | 53 | No | GU553378 |
| ACAPRb | Ppgm2 | (AACT)5 | GGTCTGCCAGGTCATTTCTC | AACGGCACTCATCTGGTTAGT | 53 | Yes | GU553379 |
| AE1 | Ppgm3 | (GA)8GG(GA)5 | AACACATGACATCACTGCAGC | ACAGGTAAGTCAGTTGTTTCAGG | 53 | Yes | GU553380 |
| AQP9 | Ppgm4 | (TGT)6 | AGAAAATAAGCAGCCGTAGC | TGCACGTAAATGGTCTGATT | 55 | No | GU553381 |
| ATP1A1 | Ppgm5 | (TG)6 | CCATAGGACGATCACAAG | GAATGAAGTCTTTGTTGTGGGTC | 53 | No | GU553382 |
| Ppgm6 | (CA)5 | AGCAGAGCAAAGAACAGGACTC | GATCTCTTTTGCTCTGGAGTTGG | 53 | No | GU553382 | |
| ATP1A2 | Ppgm7 | (TG)5/(GT)12 | TGCATAATGGTCCCCCGTG | AGGCCTTGGCATCCCTG | 53 | Yes | GU553383 |
| ATP4A | Ppgm8 | (TC)5 | TCATTGTAATTTCCGCCTTT | TTTCATCACCAACAGGTAGC | 55 | Yes | GU553384 |
| ATP6V1Aa | Ppgm9 | (TG)6/(TG)10 | GACCGATTTCATCTCTGGAC | TGACTCTTTTCCCTCCACTT | 55 | Yes | GU553385 |
| ATP6V1Ab | Ppgm10 | (CA)6 | GCAGGATACCCGCTGTCT | AAGTTCACAAAGGATGCACA | 55 | Yes | GU553386 |
| CFTR | Ppgm11 | (CA)13 | CACTGCTAACACACATCAGC | AAGCGATACCCATCTGTCC | 55 | Yes | GU553387 |
| CLCN3 | Ppgm12 | (CA)11 | AGTCGGCATGGGAGTTCAC | GCGATGTCAATCAGGCCG | 53 | Yes | GU553388 |
| CLCN4 | Ppgm13 | (CTC)5 | GTTTGAATCCCACAACTTCA | ACTACGTCAACAACCCCAAC | 55 | No | GU553389 |
| CLCN7 | Ppgm14 | (TG)10 | CGCTCTGAACAGCTTAAACA | ACGAGAGGGAGTGCATGA | 55 | Yes | GU553390 |
| CLCNK | Ppgm15 | (TA)4(CA)(TA) | GGCCACCTAGAATTGATGAC | TCAGAGTCCAAACACGAGAA | 55 | No | GU553391 |
| CSP2 | Ppgm16 | (AAAT)3 | CGTCGAACTCTACAACCTCC | AGACGTGTTTGTTCATCAGG | 55 | Yes | GU553392 |
| DIO1 | Ppgm17 | (TC)9 | CAATCAGGATGTCCAACCA | GCAGCATGGGATGAGAAC | 55 | Yes | GU553393 |
| eEF1A1b | Ppgm18 | (AC)9 | CACTACAGAGTCTAGTCTGAG | TCTTCAGTTAAATGAACCGGTTGC | 55 | Yes | GU553394 |
| FERH1 | Ppgm19 | (ATC)5 | TTCTCTCGTTTCTCCAGAGC | GAAATTGACACTGCTGGTTG | 55 | Yes | GU553395 |
| FGF6a | Ppgm20 | C12 | CATCCTTCACCCCAATCTTA | TCTGTCCCCTCTTTCAATGT | 55 | Yes | GU553396 |
| FGF18 | Ppgm21 | (AG)12 | TGCCTACTCACACCCACTAA | ATGAGAAATCAATGGAGGGA | 55 | Yes | GU553397 |
| GH | Ppgm22 | (GTATA)3 | TGCGTGGTGTAGTATAGTGTAGTC | AGAGCAACGTCAACTCAACA | 55 | No | GU553398 |
| GHRH | Ppgm23 | (AT)8 | AAGATGAGTTTCCCGCTCTA | TTATTGACTTGACCCTTGACC | 55 | No | GU553399 |
| GHR-1 | Ppgm24 | (AC)7 | CCACTACCTCTGCCCTAAAA | TTTCCTTTGGCTTCAATCTC | 55 | Yes | GU553400 |
| GHR-2 | Ppgm25 | (TA)6 | TCAACTCTGACCTTCTTGAGG | ACCACAGGTTCACCAAAGAT | 55 | Yes | GU553401 |
| GR1 | Ppgm26 | A8 | CTGGTACTGTCCTGATGGAG | TTCTCATAACCACAACTGGC | 55 | Yes | GU553402 |
| GR2 | Ppgm27 | A8 | AGAGCACGACAAAACACAGA | AGCAGAAATTGAACAGCACA | 55 | No | GU553403 |
| GTF2B | Ppgm28 | (AC)4 | TGTAATCCCAATACGACGC | AGTATCTGAACCCGCACATT | 55 | Yes | GU553404 |
| HPX | Ppgm29 | (AC)26 | GTGCTTTTAGAAAGACCACCG | TATTACTCTATAGCCGGCAGC | 55 | Yes | GU553405 |
| HSP25 | Ppgm30 | (AC)13 | GCAGCGTACATTCTGTTCAAC | GGTTTCTTATGTGGGTGTGAC | 53 | Yes | GU553406 |
| HSP47a | Ppgm31 | (CA)6 | TGCATCATCTGCACTGAAACG | GGGGCAATGATCGTCAATG | 55 | Yes | GU553407 |
| HSP70Aa | Ppgm32 | (AC)15 | CAAAGACCTGCACACACATT | GGGAGCTGTCGATACGTTTA | 55 | Yes | GU553408 |
| HSP70Ab | Ppgm33 | (AC)7 | ATCTACAGGGATACCACAGTAC | TGTTTACTCCGGTCAATGAAACC | 53 | No | GU553409 |
| HSP70Ac | Ppgm34 | (TG)5 | TACTGTTCCACTTGCCCATT | TCACAACTCAGGATCTCGAA | 55 | No | GU553410 |
| HSP70B | Ppgm35 | (TC)14 | TGAGGGTAAAAGCTGTAGCA | ATTATCCCAGAACACTCCCA | 55 | Yes | GU553411 |
| HSP90Ab | Ppgm36 | (CA)5 | GTCAAACCGGACATTAGGAC | CAGACGTGAAACTACGCTTG | 55 | Yes | GU553412 |
| HSP90B | Ppgm37 | (TCG)4 | AGTTATGAAGAAACCGCGTC | GTGATGGCTGTAGCTTGTTG | 55 | Yes | GU553413 |
| IGF-I | Ppgm38 | (CGC)4 | AAGGACGAGCTCGGCTAC | AGGATGCGGCTGCAGATG | 53 | No | GU553414 |
| IGF-II | Ppgm39 | (TA)13/(TA)5 | GTTAGGCTTTTACTTGGGTTTCC | TCATTACGCAAGATACAGCTCAG | 53 | Yes | GU553415 |
| Kir2.1a | Ppgm40 | (TA)10 | CTGGAATGACACCAGCCAG | AGTCAGCTCCAAGCTTGTG | 53 | Yes | GU553416 |
| Kir2.1c | Ppgm41 | (TG)7 | CAAAGTGAGCAACAGTGAGC | AAAGGTCGAGGAAGATGATG | 55 | No | GU553417 |
| Kir2.2 | Ppgm42 | (TC)12 | ACAGGTACGAAGCGTTTAGC | AGAGCAAAGAAATAGACGGG | 55 | Yes | GU553418 |
| MSTNa | Ppgm43 | (CA)5 | AGTCCGCAATAAGCTCAAAC | ACAAAGCCGTCTAGGTGTTC | 55 | No | GU553419 |
| MSTNb | Ppgm44 | (GGA)8 | GCTGGAGCAGTACGACC | GGTGATGATGGTCTCCGT | 55 | No | GU553420 |
| MYHa | Ppgm45 | (TA)3C(TA)3 | AAAGCTCCAAGTAACGCTGT | ACTTTTGTTTCCAATCTGCC | 55 | No | GU553421 |
| MYHe | Ppgm46 | (GTAA)4 | ATGTTAATTGCTTTGTGCGA | TGAAACACAGGAGCTTGAGA | 55 | Yes | GU553422 |
| NKCC1b | Ppgm47 | (CA)13 | ATATGTTCAGCACGCAGCG | GTCAAAGGAGTCTTAGTGAGTG | 55 | Yes | GU553423 |
| NPY2Rb | Ppgm48 | (GT)26 | CACTCAGGAGAAGTGAGGC | GGAACGATTACAGGTACGGAC | 53 | Yes | GU553424 |
| NPYP | Ppgm49 | T10 | CTGCAGCGGACGGGATTAG | AACACAGGACCGACTTTGAGG | 53 | No | GU553425 |
| PKMa | Ppgm50 | (GA)5 | GTGGTACTGCTGGTTGTACT | AAGGTCAAACGGCGTCGC | 53 | Yes | GU553426 |
| PVALBb | Ppgm51 | (CA)6 | AGCTGAACTTTGGTGTGTCTC | GTTGATGTGCATTTATGGGA | 55 | Yes | GU553427 |
| SHH | Ppgm52 | G18 | AGAAACGTGGTTATTGAGGC | CCTGCTCTTTATTGGGTTTT | 55 | Yes | GU553428 |
| SLC14 | Ppgm53 | (TG)13 | CCATCATCTCTACACATAATCAAAC | CACTGATTACAGATTGTGTGCTG | 53 | Yes | GU553429 |
| SSR1b | Ppgm54 | (TG)12 | TGTCCTCGCAAAGTTCATAA | TGACCGAGCATTTTACTTGA | 55 | Yes | GU553430 |
| T1R3 | Ppgm55 | T18 | ATGCACTGCGTTATCACTCC | GCTTTGTTTAACGTCAGTATTTCG | 55 | Yes | GU553431 |
| TAAR | Ppgm56 | (AC)12 | CAAGGACCACGCTAAAGGTA | GATTTCTTCTTGATGTCCGC | 55 | Yes | GU553432 |
| TBX4 | Ppgm57 | (GT)8 | GAGTACGAGCAGGTTTGGTT | GTAACACACACGGTTTTGGA | 55 | Yes | GU553433 |
| TTP | Ppgm58 | (AC)3(TC)(AC)3 | GCCACTTATACATGCTCGC | GATTCTCCCTGCCTCACA | 55 | Yes | GU553434 |
Ta, annealing temperature.
Table 3.
Genetic variability of SSR markers for functionally important genes in three populations of nine-spined sticklebacks.
| Gene ID | Locus | Total A | Baltic Sea | Lake 1 | Pyöreälampi | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Size (bp) | A | HE | FIS | Size (bp) | A | HE | FIS | Size (bp) | A | HE | FIS | |||
| ACAPRb | Ppgm2 | 3 | 382-414 | 3 | 0.573 | -0.017 | 410 | 1 | 0.000 | na | 414 | 1 | 0.000 | na |
| AE1 | Ppgm3 | 15 | 214-260 | 10 | 0.779 | -0.070 | 226-308 | 6 | 0.560 | 0.146 | 228 | 1 | 0.000 | na |
| ATP1A2 | Ppgm7 | 9 | 117-143 | 9 | 0.774 | -0.077 | 117-139 | 3 | 0.576 | -0.085 | 135 | 1 | 0.000 | na |
| ATP4A | Ppgm8 | 2 | 186-196 | 2 | 0.042 | 0.000 | 186 | 1 | 0.000 | na | 186 | 1 | 0.000 | na |
| ATP6V1Aa | Ppgm9 | 7 | 187-199 | 7 | 0.783 | -0.065 | 189-191 | 2 | 0.504 | -0.324 | 191 | 1 | 0.000 | na |
| ATP6V1Ab | Ppgm10 | 4 | 102-110 | 3 | 0.265 | 0.058 | 98-110 | 2 | 0.451 | -0.293 | 110 | 1 | 0.000 | na |
| CFTR | Ppgm11 | 10 | 129-159 | 9 | 0.593 | 0.157 | 139-143 | 2 | 0.359 | -0.278 | 149 | 1 | 0.000 | na |
| CLCN3 | Ppgm12 | 25 | 185-241 | 20 | 0.935 | -0.023 | 194-225 | 6 | 0.755 | 0.118 | 193-195 | 2 | 0.437 | -0.241 |
| CLCN7 | Ppgm14 | 8 | 245-266 | 6 | 0.626 | 0.001 | 254-261 | 4 | 0.746 | 0.107 | 256 | 1 | 0.000 | na |
| CSP2 | Ppgm16 | 3 | 184-192 | 3 | 0.490 | -0.360 | 184 | 1 | 0.000 | na | 184 | 1 | 0.000 | na |
| DIO1 | Ppgm17 | 4 | 162-166 | 4 | 0.545 | 0.123 | 162-166 | 2 | 0.254 | -0.150 | 166 | 1 | 0.000 | na |
| eEF1A1b | Ppgm18 | 5 | 217-229 | 5 | 0.715 | 0.125 | 227-229 | 2 | 0.223 | -0.122 | 225 | 1 | 0.000 | na |
| FERH1 | Ppgm19 | 4 | 217-235 | 4 | 0.352 | 0.054 | 232-235 | 2 | 0.496 | -0.007 | 235 | 1 | 0.000 | na |
| FGF6a | Ppgm20 | 14 | 332-346 | 12 | 0.859 | -0.063 | 335-340 | 4 | 0.624 | 0.132 | 337 | 1 | 0.000 | na |
| FGF18 | Ppgm21 | 6 | 331-347 | 6 | 0.723 | -0.038 | 335-343 | 2 | 0.511 | 0.021 | 335 | 1 | 0.000 | na |
| GHR-1 | Ppgm24 | 5 | 193-205 | 3 | 0.121 | -0.030 | 192-193 | 2 | 0.223 | -0.122 | 193-195 | 2 | 0.156 | -0.070 |
| GHR-2 | Ppgm25 | 3 | 327-333 | 3 | 0.492 | -0.186 | 331-333 | 2 | 0.082 | -0.022 | 327 | 1 | 0.000 | na |
| GR1 | Ppgm26 | 3 | 295-297 | 3 | 0.532 | 0.060 | 297 | 1 | 0.000 | na | 297 | 1 | 0.000 | na |
| GTF2B | Ppgm28 | 2 | 290 | 1 | 0.000 | na | 288-290 | 2 | 0.418 | -0.394 | 290 | 1 | 0.000 | na |
| HPX | Ppgm29 | 13 | 126-169 | 10 | 0.746 | 0.163 | 137-157 | 3 | 0.586 | 0.289 | 155-158 | 2 | 0.504 | 0.090 |
| HSP25 | Ppgm30 | 4 | 159-177 | 4 | 0.605 | 0.243 | 159-175 | 2 | 0.478 | -0.045 | 173 | 1 | 0.000 | na |
| HSP47a | Ppgm31 | 2 | 107-109 | 2 | 0.042 | 0.000 | 107 | 1 | 0.000 | na | 107 | 1 | 0.000 | na |
| HSP70Aa | Ppgm32 | 27 | 283-529 | 26 | 0.959 | 0.094 | 305-329 | 4 | 0.662 | 0.146 | 317-319 | 2 | 0.466 | -0.163 |
| HSP70B | Ppgm35 | 6 | 324-348 | 6 | 0.416 | -0.102 | 324-342 | 2 | 0.462 | -0.533 | 324-342 | 2 | 0.283 | -0.179 |
| HSP90Ab | Ppgm36 | 7 | 198-232 | 7 | 0.629 | 0.205 | 204-206 | 2 | 0.156 | -0.070 | 204 | 1 | 0.000 | na |
| HSP90B | Ppgm37 | 7 | 244-271 | 6 | 0.728 | 0.085 | 232-259 | 3 | 0.429 | -0.068 | 247 | 1 | 0.000 | na |
| IGF-II | Ppgm39 | 8 | 387-401 | 8 | 0.700 | 0.107 | 389-397 | 3 | 0.160 | 0.480 | 393-397 | 3 | 0.159 | -0.051 |
| Kir2.1a | Ppgm40 | 8 | 293-327 | 8 | 0.713 | -0.169 | 301-321 | 2 | 0.511 | 0.103 | 297-301 | 2 | 0.156 | -0.070 |
| Kir2.2 | Ppgm42 | 6 | 289-299 | 6 | 0.578 | -0.082 | 291-293 | 2 | 0.223 | -0.122 | 289 | 1 | 0.000 | na |
| MYHe | Ppgm46 | 6 | 154-161 | 5 | 0.639 | 0.021 | 154-159 | 3 | 0.508 | -0.148 | 159 | 1 | 0.000 | na |
| NKCC1b | Ppgm47 | 10 | 117-159 | 10 | 0.744 | -0.065 | 119-159 | 3 | 0.629 | 0.171 | 123-125 | 2 | 0.042 | 0.000 |
| NPY2Rb | Ppgm48 | 25 | 406-471 | 19 | 0.918 | 0.001 | 423-475 | 6 | 0.793 | 0.002 | 469 | 1 | 0.000 | na |
| PKMa | Ppgm50 | 5 | 327-334 | 3 | 0.518 | 0.660 | 327-334 | 3 | 0.392 | -0.062 | 327-329 | 3 | 0.159 | -0.045 |
| PVALBb | Ppgm51 | 5 | 168-176 | 5 | 0.747 | 0.243 | 169-176 | 2 | 0.042 | 0.000 | 169 | 1 | 0.000 | na |
| SHH | Ppgm52 | 13 | 344-356 | 11 | 0.856 | 0.054 | 353-358 | 2 | 0.315 | 0.603 | 351-353 | 2 | 0.198 | -0.100 |
| SLC14 | Ppgm53 | 3 | 230-240 | 3 | 0.121 | -0.030 | 236 | 1 | 0.000 | na | 236 | 1 | 0.000 | na |
| SSR1b | Ppgm54 | 8 | 129-145 | 8 | 0.544 | 0.005 | 135-145 | 4 | 0.727 | 0.083 | 139 | 1 | 0.000 | na |
| T1R3 | Ppgm55 | 6 | 278-285 | 5 | 0.558 | -0.195 | 281-286 | 2 | 0.082 | -0.022 | 281 | 1 | 0.000 | na |
| TAAR | Ppgm56 | 9 | 123-141 | 8 | 0.614 | 0.050 | 123-139 | 4 | 0.659 | 0.305 | 127 | 1 | 0.000 | na |
| TBX4 | Ppgm57 | 3 | 278-292 | 3 | 0.318 | -0.048 | 278-290 | 2 | 0.380 | -0.314 | 290 | 1 | 0.000 | na |
| TTP | Ppgm58 | 3 | 317-321 | 3 | 0.291 | 0.140 | 317-321 | 2 | 0.283 | -0.179 | 317 | 1 | 0.000 | na |
A, number of observed alleles; HE, expected heterozygosity; na, not applied.
Patterns and degree of SSR variability
The level of SSR variability is known to be associated with repeat motifs due to their different mutation rates [61,62]. In addition, cross-species transfer of SSR primers often results in a lower level of SSR variability in a focal species relative to a source species because of ascertainment bias [63,64]. We investigated the patterns and degree of SSR variability using three Fennoscandian populations. Across the 104 polymorphic loci identified in this study, an average number of alleles per locus and average heterozygosity were 7.4 and 0.60 in the Baltic Sea, 2.6 and 0.32 in the Lake 1, and 1.6 and 0.10 in the Pyöreälampi, respectively. As expected, the levels of SSR variability were significantly dependent on population. The genome-wide survey indicated that genetic variation of the Pyöreälampi is very low, as also shown in a previous study with 11 SSR and one insertion/deletion loci [65]. In our data set, the levels of SSR variability were not dependent on marker origin, SSR type and SSR repeat motif (Table 4). However, a significant influence of SSR location on the levels of allele number and heterozygosity was apparent (Table 4). Across the three populations, average allele number and heterozygosity were 4.7 and 0.41 in exonic regions, 3.3 and 0.30 in intronic regions and 4.2 and 0.36 in intergenic regions, respectively. While the level of SSR variability is known to differ between coding and untranslated regions [66,67], EST-derived SSRs tend to show lower variability than genomic SSRs [50,68]. These differences are thought to arise due to heterogeneous distributions of SSR repeat motifs. However, the higher variability in exonic SSRs than in other SSRs is not explainable by an artifact stemming from different repeat motifs because a majority of the polymorphic SSRs were dimeric repeats independently of their location. Several lines of evidence suggest that SSR variation may affect various traits and be subject to natural selection [21,69,70]. While the potential effect of variable mutation rates can not be ruled out, the heterogeneous distribution of SSR variability observed in this study might be ascribable to natural selection.
Table 4.
Hierarchical analysis of genetic variability in nine-spined sticklebacks.
| Source | Number of alleles | Expected heterozygosity | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Type III SS | df (den) | MS | F | P | Type III SS | df (den) | MS | F | P | |
| Marker origin | 2.653 | 1 (298) | 2.653 | 0.222 | 0.638 | 0.029 | 1 (298) | 0.029 | 0.527 | 0.468 |
| SSR type | 16.859 | 2 (298) | 8.430 | 0.704 | 0.495 | 0.079 | 2 (298) | 0.040 | 0.716 | 0.489 |
| SSR location | 73.246 | 2 (298) | 36.623 | 3.059 | 0.048 | 0.416 | 2 (298) | 0.208 | 3.782 | 0.024 |
| SSR motif | 36.085 | 3 (298) | 12.028 | 1.005 | 0.391 | 0.073 | 3 (298) | 0.024 | 0.445 | 0.721 |
| Population | 1978.000 | 2 (298) | 989.000 | 12.587 | 2 (298) | 6.294 | ||||
Conclusions
Our study demonstrated that a large proportion of SSRs are conserved in the stickleback species which have diverged from a common ancestor more than 10 million years ago [46]. Therefore, the three-spined stickleback genome can be used to predict SSR locations in Pungitius species. Our results also suggest that the main limitation of cross-species utility of SSR markers lies in the failure of amplification success probably due to mutations in SSR flanking sequences. While it is possible to predict to some degree the likelihood of amplification success based on the information of primer binding sites, cross-species transferability of SSR primers for functionally important genes is particularly low as compared to that of genomic and EST-derived SSR primers. Yet, SSR markers can be developed for functionally important genes and target genomic regions using the approach outlined in this paper. This approach should be applicable also to other non-model organisms. The SSR markers developed for functionally important genes should be useful to identify genes responsible for phenotypic variation and adaptive divergence in nine-spined sticklebacks, as well as for constructing comparative gene maps of nine-spined and three-spined sticklebacks.
Methods
Fish samples
Nine-spined sticklebacks collected from the Baltic Sea (coastal; 60°12' N, 25°11' E), the 'Lake 1' (lake; 67°54' N, 20°50' E) and the Pyöreälampi (pond; 66°16' N, 29°26' E) were used in this study. The fish were sampled with seine nets or minnow traps in 2002 (Lake 1) and 2008 (Baltic Sea and Pyöreälampi). Total DNA was extracted from fin clips stored in 70-99% ethanol with a phenol-chloroform method [71] following proteinase K digestion.
Cross-species transfer of three-spined stickleback SSR primers
Cross-species utility of three-spined stickleback SSR primers was tested for 158 SSR markers for physiologically important genes (gene-based SSRs) [Y. Shimada, T. Shikano and J. Merilä, unpublished] coupled with 101 markers derived from genomic libraries (genomic SSRs) and 87 markers derived from ESTs (EST-derived SSRs; Additional files 1 and 2) [72-76]. The genomic and EST-derived SSRs were classified according to the source information deposited in GenBank [77]. The following factors potentially affecting cross-species amplification success were scored for each of the makers: SSR marker type (cf. gene-based, genomic and EST-derived SSRs), primer binding site (cf. exonic, intronic, intergenic and other combinations), average primer length, average and difference of GC content and melting temperature in forward and reverse primer pairs, as well as expected PCR product size. Primer binding sites were categorized into exonic, intronic and intergenic regions based on the Ensembl genebuild in the three-spined stickleback genome [40]. Since information on untranslated regions was not available for a number of genes, we did not distinguish between coding and untranslated regions in the analyses. The primer parameters were calculated using BioEdit [78] under the actual PCR conditions (see below). The expected PCR product sizes were calculated based on the three-spined stickleback genome. The role of these factors was evaluated using generalized linear models as implemented in JMP 5 (SAS Inst. Inc.). In these tests, amplification success was treated as a binary dependent variable (successful amplification = 1, failed amplification = 0), SSR type and primer site as factors, and other parameters as covariates. Logit link function was used. For the successfully amplified loci, factors affecting incidence of polymorphism were evaluated using generalized linear models treating SSR type, SSR location (cf. exonic, intronic and intergenic regions) and SSR repeat motif (cf. di- and trinucleotide repeats) as factors. SSR location was categorized into exonic, intronic and intergenic regions based on the Ensembl genebuild in the three-spined stickleback genome. In this test, polymorphic locus was treated as a binary dependent variable (polymorphic = 1, monomorphic = 0) using logit link function.
SSR primer development in nine-spined sticklebacks
Based on the literature on gene functions in teleosts, we selected 67 genes responsible for significant physiological - such as osmoregulation, thermal response, growth, disease and taste [Y. Shimada, T. Shikano and J. Merilä, unpublished] - and developmental functions [e.g. [41]] (Additional file 3). Genomic locations of these genes were identified in the three-spined stickleback genome following Shimada et al. [Y. Shimada, T. Shikano and J. Merilä, unpublished]. In brief, we searched the three-spined stickleback ESTs which correspond to target genes of this species or other teleosts in the GenBank database [77] and mapped them in the three-spined stickleback genome. The genomic range of respective genes was determined according to the Ensembl transcript and Genscan predictions (Additional file 3). Since the genomic region of the PITX1 was not available due to partially incomplete sequences of the three-spined stickleback genome, the sequence of this gene (GenBank: AY517634.1) was used.
SSRs were searched in the target genes and their flanking regions in the three-spined stickleback genome using Tandem repeats finder [79]. In order to survey conserved regions for designing amplification and sequencing primers in nine-spined sticklebacks, the sequences of these genomic regions were subject to BLASTN searches against the currently available genome sequences of other teleosts, i.e. medaka, fugu, spotted green pufferfish and/or zebrafish [40]. Conserved regions were determined by aligning the sequences of three-spined sticklebacks and those of other fish species detected by the BLASTN searches. Based on the location of SSRs and conserved regions, primer sequences for nine-spined sticklebacks were designed manually in one genomic region for each target gene, except for the GHRI and IGF-I, for which two and three regions were used, respectively (Additional file 3).
Two individuals of the Pyöreälampi were used for amplifying and sequencing the target genomic regions. One three-spined stickleback individual from the Baltic Sea (60°12' N, 25°11' E) was used as a positive control. Using a primer pair for respective target regions (Additional file 3), PCR amplifications were carried out in a 20 μl reaction volume consisting of 1× PCR buffer (Bioline), 1.5 mM MgCl2, 0.25 mM dNTP (Finnzymes), 0.15 U BIOTAQ DNA polymerase (Bioline), 5 pmol of each primer and approx. 40 ng of genomic DNA. The reactions were performed as follows: an initial degeneration step at 95°C for 3 min, followed by 30 s at 95°C, 30 s at 53-60°C and 60-120 s at 72°C for 35 cycles with a final extension at 72°C for 5 min (see Additional file 3 for optimal PCR conditions in each primer pair). Approximate size of the PCR amplicons was determined by electrophoresis on 1.5% agarose gel with a DNA ladder (GeneRuler™ DNA Ladder Mix, Fermentas). PCR products were purified using exonuclease I (New England Biolabs) and shrimp alkaline phosphatase (Roche) and directly sequenced in both forward and reverse directions with the same primers as those used in the PCRs. The sequencing reactions were performed using the BigDye Terminator v3.1 Cycle Sequencing Kit (Applied Biosystems) according to manufacture's instructions. Cycle sequencing products were purified by ethanol precipitation and analyzed on an ABI 3730xl DNA Analyzer (Applied Biosystems).
The sequences in forward and reverse directions of two individuals were aligned using CLUSTAL W [80] as implemented in MEGA 4 [81] and edited by hand. For large PCR amplicons (≥1200 bp), the sequences in forward and reverse directions were separately aligned using two individuals (Additional file 4). As sequences were available only in one direction for four genomic regions even after retrials, sequences for these regions were aligned using two individuals (Additional file 4). The sequences were subject to BLASTN searches against the three-spined stickleback genome to ensure that they are mapped back to the correct locations in the genome. The homologous sequences in three-spined and nine-spined sticklebacks were aligned to compare SSR locations and motifs between them. This comparison was performed using SSRs with minimum repeat numbers of ten, five and four for one (mono-), two (di-) and three or longer (tri-, tetra-, penta- and hexanucleotide) repeat motifs, respectively. To further address SSR conservation in stickleback species, we also investigated if SSRs randomly derived from genomic libraries of Pungitius species are found at the homologous locations in the three-spined stickleback genome using publicly available SSR and flanking sequences of Pungitius pungitius (i.e. nine-spined stickleback) [[56]; GenBank: AB473819-AB473831] and Pungitius sp. (Omono type) [[57]; GenBank: AB300827-AB300851]. Out of the 38 sequences, six (GenBank: AB300830, AB300831, AB300841, AB300842, AB300844, AB300849) were excluded from the analyses because of low BLAST hit scores and alignment problems. To develop SSR markers for functionally important genes, primer sets were designed based on the sequences of nine-spined sticklebacks using WebSat [82]. Primer sequences were deposited in GenBank under accession numbers GU553378-GU553434.
SSR amplification and genotyping
For the cross-species amplification test of three-spined stickleback primers, amplification success and polymorphism were determined using the following three-step procedure. Firstly, amplification was tested using four individuals from the Pyöreälampi and Baltic Sea (two individuals per population) with fluorescent labelled forward primers (FAM, HEX or TET) and GTTT-tailed reverse primers [83]. As a positive control, one individual of the three-spined stickleback was used. PCRs were performed under optimal conditions for three-spined sticklebacks and conducted in a 10 μl reaction volume consisting of 1× PCR buffer (Bioline), 1.5 mM MgCl2, 0.2 mM dNTP (Finnzymes), 0.18 U BIOTAQ DNA polymerase (Bioline), 5 pmol of each primer and approx. 20 ng of template DNA. The reactions were performed as follows: an initial degeneration step at 95°C for 3 min, followed by 30 s at 95°C, 30 s at 53°C and 30 s at 72°C for 30 cycles with a final extension at 72°C for 5 min. Amplification success was determined by electrophoresis on 1.6% agarose gel. Secondly, for the loci that showed robust and specific amplification within the expected size range, polymorphism was investigated by genotyping 24 individuals from the Baltic Sea and the Lake 1 (12 individuals per population). For efficient screening, PCRs were carried out using the Qiagen Multiplex PCR Kit (Qiagen) in 10 μl reaction volumes containing 1× Qiagen Multiplex PCR Master Mix, 0.5× Q-Solution, 2 pmol of each primer and approx. 20 ng of template DNA. The reactions were performed by the following cycle: an initial activation step at 95°C for 15 min, followed by 30 s at 94°C, 90 s at 53°C and 60 s at 72°C for 30 cycles with a final extension at 60°C for 5 min. PCR products were visualized with a MegaBACE 1000 automated sequencer (Amersham Biosciences) and their sizes were determined with ET-ROX 550 size standard (Amersham Biosciences). Thirdly, genetic variability of the polymorphic loci identified with the 24 individuals was evaluated by genotyping a total of 24 individuals from each of the Baltic Sea, Lake 1 and Pyöreälampi populations using multiplex PCRs. For the SSR primers developed in nine-spined sticklebacks, polymorphism and genetic variability were evaluated following the procedures for the second and third steps. Since some of the SSR markers yielded clearer allele profiles at an annealing temperature of 55°C, this temperature was used for these loci instead of 53°C (see Table 2 for an optimal annealing temperature for each primer pair). Alleles were scored using Fragment Profiler 1.2 (Amersham Biosciences) with visual inspection and manual corrections of alleles.
SSR data analyses
Locus and population specific gene diversities (HE) [84] were estimated using FSTAT 2.9.3 [85,86]. Within population and locus specific FIS were estimated for each population to detect possible deviations from Hardy-Weinberg equilibrium with 10 000 permutations using FSTAT 2.9.3. Sequential Bonferroni corrections [87] were applied to minimize type I errors. The presence of null alleles was tested using MICRO-CHECKER [88].
Factors affecting the levels of genetic variation were evaluated with general linear models where allele number or heterozygosity was treated as a dependent variable, SSR marker origin (cf. three-spined and nine-spined sticklebacks), SSR type, SSR location and SSR repeat motif (cf. mono-, di-, tri- and tetranucleotide repeats) as fixed factors and population as a random factor. These analyses were performed with JMP 5.
Authors' contributions
TS conceived the study, contributed to the gene selection and localization, prepared the molecular data, conducted the analyses and wrote the manuscript. JR selected the genes and conducted the molecular work. YS made contributions to the gene selection and localization. JM advised on the statistical analyses and contributed to writing the manuscript. All authors read and approved the final manuscript.
Supplementary Material
Genetic variability of three-spined stickleback SSR markers in three populations of nine-spined sticklebacks.
List of three-spined stickleback SSR markers that were not polymorphic or successfully amplified in nine-spined sticklebacks.
Target genes and their function and location in the theree-spined stickleback genome, and PCR and sequencing primers for nine-spined sticklebacks.
Homology of nine-spined stickleback sequences in the three-spined stickleback genome and comparative genomic location of SSRs in these species.
Contributor Information
Takahito Shikano, Email: takahito.shikano@helsinki.fi.
Jetty Ramadevi, Email: jramadevi@gmail.com.
Yukinori Shimada, Email: yukinori.shimada@helsinki.fi.
Juha Merilä, Email: juha.merila@helsinki.fi.
Acknowledgements
We thank people at the Oulanka Research Station, Abigel Gonda, Aki Hirvonen, John Loehr and Jarmo Saarikivi for help in obtaining samples. Thanks are also due to Craig Primmer for comments and checking the English. Our study was supported by the Academy of Finland and the Japan Society for the Promotion of Science.
References
- Aerts S, Lambrechts D, Maity S, Van Loo P, Coessens B, De Smet F, Tranchevent LC, De Moor B, Marynen P, Hassan B, Carmeliet P, Moreau Y. Gene prioritization through genomic data fusion. Nat Biotechnol. 2006;24:537–544. doi: 10.1038/nbt1203. [DOI] [PubMed] [Google Scholar]
- Peña-Castillo L, Tasan M, Myers CL, Lee H, Joshi T, Zhang C, Guan Y, Leone M, Pagnani A, Kim WK, Krumpelman C, Tian W, Obozinski G, Qi Y, Mostafavi S, Lin GN, Berriz GF, Gibbons FD, Lanckriet G, Qiu J, Grant C, Barutcuoglu Z, Hill DP, Warde-Farley D, Grouios C, Ray D, Blake JA, Deng M, Jordan MI, Noble WS, Morris Q, Klein-Seetharaman J, Bar-Joseph Z, Chen T, Sun F, Troyanskaya OG, Marcotte EM, Xu D, Hughes TR, Roth FP. A critical assessment of Mus musculus gene function prediction using integrated genomic evidence. Genome Biol. 2008;9:52. doi: 10.1186/gb-2008-9-s1-s2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann AA, Willi Y. Detecting genetic responses to environmental change. Nat Rev Genet. 2008;9:421–432. doi: 10.1038/nrg2339. [DOI] [PubMed] [Google Scholar]
- Bonin A. Population genomics: a new generation of genome scans to bridge the gap with functional genomics. Mol Ecol. 2008;17:3583–3584. doi: 10.1111/j.1365-294X.2008.03854.x. [DOI] [PubMed] [Google Scholar]
- Nielsen EE, Hemmer-Hansen J, Larsen PF, Bekkevold D. Population genomics of marine fishes: identifying adaptive variation in space and time. Mol Ecol. 2009;18:3128–3150. doi: 10.1111/j.1365-294X.2009.04272.x. [DOI] [PubMed] [Google Scholar]
- O'Brien SJ, Womack JE, Lyons LA, Moore KJ, Jenkins NA, Copeland NG. Anchored reference loci for comparative genome mapping in mammals. Nat Genet. 1993;3:103–112. doi: 10.1038/ng0293-103. [DOI] [PubMed] [Google Scholar]
- Lyons LA, Laughlin TF, Copeland NG, Jenkins NA, Womack JE, O'Brien SJ. Comparative anchor tagged sequences (CATS) for integrative mapping of mammalian genomes. Nat Genet. 1997;15:47–56. doi: 10.1038/ng0197-47. [DOI] [PubMed] [Google Scholar]
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J, Devon K, Dewar K, Doyle M, FitzHugh W, Funke R, Gage D, Harris K, Heaford A, Howland J, Kann L, Lehoczky J, LeVine R, McEwan P, McKernan K, Meldrim J, Mesirov JP, Miranda C, Morris W, Naylor J, Raymond C, Rosetti M, Santos R, Sheridan A, Sougnez C, Stange-Thomann N, Stojanovic N, Subramanian A, Wyman D, Rogers J, Sulston J, Ainscough R, Beck S, Bentley D, Burton J, Clee C, Carter N, Coulson A, Deadman R, Deloukas P, Dunham A, Dunham I, Durbin R, French L, Grafham D, Gregory S, Hubbard T, Humphray S, Hunt A, Jones M, Lloyd C, McMurray A, Matthews L, Mercer S, Milne S, Mullikin JC, Mungall A, Plumb R, Ross M, Shownkeen R, Sims S, Waterston RH, Wilson RK, Hillier LW, McPherson JD, Marra MA, Mardis ER, Fulton LA, Chinwalla AT, Pepin KH, Gish WR, Chissoe SL, Wendl MC, Delehaunty KD, Miner TL, Delehaunty A, Kramer JB, Cook LL, Fulton RS, Johnson DL, Minx PJ, Clifton SW, Hawkins T, Branscomb E, Predki P, Richardson P, Wenning S, Slezak T, Doggett N, Cheng JF, Olsen A, Lucas S, Elkin C, Uberbacher E, Frazier M, Gibbs RA, Muzny DM, Scherer SE, Bouck JB, Sodergren EJ, Worley KC, Rives CM, Gorrell JH, Metzker ML, Naylor SL, Kucherlapati RS, Nelson DL, Weinstock GM, Sakaki Y, Fujiyama A, Hattori M, Yada T, Toyoda A, Itoh T, Kawagoe C, Watanabe H, Totoki Y, Taylor T, Weissenbach J, Heilig R, Saurin W, Artiguenave F, Brottier P, Bruls T, Pelletier E, Robert C, Wincker P, Smith DR, Doucette-Stamm L, Rubenfield M, Weinstock K, Lee HM, Dubois J, Rosenthal A, Platzer M, Nyakatura G, Taudien S, Rump A, Yang H, Yu J, Wang J, Huang G, Gu J, Hood L, Rowen L, Madan A, Qin S, Davis RW, Federspiel NA, Abola AP, Proctor MJ, Myers RM, Schmutz J, Dickson M, Grimwood J, Cox DR, Olson MV, Kaul R, Raymond C, Shimizu N, Kawasaki K, Minoshima S, Evans GA, Athanasiou M, Schultz R, Roe BA, Chen F, Pan H, Ramser J, Lehrach H, Reinhardt R, McCombie WR, de la Bastide M, Dedhia N, Blöcker H, Hornischer K, Nordsiek G, Agarwala R, Aravind L, Bailey JA, Bateman A, Batzoglou S, Birney E, Bork P, Brown DG, Burge CB, Cerutti L, Chen HC, Church D, Clamp M, Copley RR, Doerks T, Eddy SR, Eichler EE, Furey TS, Galagan J, Gilbert JG, Harmon C, Hayashizaki Y, Haussler D, Hermjakob H, Hokamp K, Jang W, Johnson LS, Jones TA, Kasif S, Kaspryzk A, Kennedy S, Kent WJ, Kitts P, Koonin EV, Korf I, Kulp D, Lancet D, Lowe TM, McLysaght A, Mikkelsen T, Moran JV, Mulder N, Pollara VJ, Ponting CP, Schuler G, Schultz J, Slater G, Smit AF, Stupka E, Szustakowski J, Thierry-Mieg D, Thierry-Mieg J, Wagner L, Wallis J, Wheeler R, Williams A, Wolf YI, Wolfe KH, Yang SP, Yeh RF, Collins F, Guyer MS, Peterson J, Felsenfeld A, Wetterstrand KA, Patrinos A, Morgan MJ, de Jong P, Catanese JJ, Osoegawa K, Shizuya H, Choi S, Chen YJ. Initial sequencing and analysis of the human genome. Nature. 2001;409:860–921. doi: 10.1038/35057062. [DOI] [PubMed] [Google Scholar]
- Warren WC, Hillier LW, Marshall Graves JA, Birney E, Ponting CP, Grützner F, Belov K, Miller W, Clarke L, Chinwalla AT, Yang SP, Heger A, Locke DP, Miethke P, Waters PD, Veyrunes F, Fulton L, Fulton B, Graves T, Wallis J, Puente XS, López-Otín C, Ordóñez GR, Eichler EE, Chen L, Cheng Z, Deakin JE, Alsop A, Thompson K, Kirby P, Papenfuss AT, Wakefield MJ, Olender T, Lancet D, Huttley GA, Smit AF, Pask A, Temple-Smith P, Batzer MA, Walker JA, Konkel MK, Harris RS, Whittington CM, Wong ES, Gemmell NJ, Buschiazzo E, Vargas Jentzsch IM, Merkel A, Schmitz J, Zemann A, Churakov G, Kriegs JO, Brosius J, Murchison EP, Sachidanandam R, Smith C, Hannon GJ, Tsend-Ayush E, McMillan D, Attenborough R, Rens W, Ferguson-Smith M, Lefèvre CM, Sharp JA, Nicholas KR, Ray DA, Kube M, Reinhardt R, Pringle TH, Taylor J, Jones RC, Nixon B, Dacheux JL, Niwa H, Sekita Y, Huang X, Stark A, Kheradpour P, Kellis M, Flicek P, Chen Y, Webber C, Hardison R, Nelson J, Hallsworth-Pepin K, Delehaunty K, Markovic C, Minx P, Feng Y, Kremitzki C, Mitreva M, Glasscock J, Wylie T, Wohldmann P, Thiru P, Nhan MN, Pohl CS, Smith SM, Hou S, Nefedov M, de Jong PJ, Renfree MB, Mardis ER, Wilson RK. Genome analysis of the platypus reveals unique signatures of evolution. Nature. 2008;453:175–183. doi: 10.1038/nature06936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zane L, Bargelloni L, Patarnello T. Strategies for microsatellite isolation: a review. Mol Ecol. 2002;11:1–16. doi: 10.1046/j.0962-1083.2001.01418.x. [DOI] [PubMed] [Google Scholar]
- Bouck A, Vision TJ. The molecular ecologist's guide to expressed sequence tags. Mol Ecol. 2007;16:907–924. doi: 10.1111/j.1365-294X.2006.03195.x. [DOI] [PubMed] [Google Scholar]
- Barbará T, Palma-Silva C, Paggi GM, Bered F, Fay MF, Lexer C. Cross-species transfer of nuclear microsatellite markers: potential and limitations. Mol Ecol. 2007;16:3759–3767. doi: 10.1111/j.1365-294X.2007.03439.x. [DOI] [PubMed] [Google Scholar]
- Cordeiro GM, Casu R, McIntyre CL, Manners JM, Henry RJ. Microsatellite markers from sugarcane (Saccharum spp.) ESTs cross transferable to erianthus and sorghum. Plant Sci. 2001;160:1115–1123. doi: 10.1016/S0168-9452(01)00365-X. [DOI] [PubMed] [Google Scholar]
- Decroocq V, Favé MG, Hagen L, Bordenave L, Decroocq S. Development and transferability of apricot and grape EST microsatellite markers across taxa. Theor Appl Genet. 2003;106:912–922. doi: 10.1007/s00122-002-1158-z. [DOI] [PubMed] [Google Scholar]
- Varshney RK, Graner A, Sorrells ME. Genic microsatellite markers in plants: features and applications. Trends Biotechnol. 2005;23:48–55. doi: 10.1016/j.tibtech.2004.11.005. [DOI] [PubMed] [Google Scholar]
- Vasemägi A, Nilsson J, Primmer CR. Seventy five EST-linked Atlantic salmon (Salmo salar L.) microsatellite markers and their cross-species amplification in salmonids. Mol Ecol Notes. 2005;5:282–288. doi: 10.1111/j.1471-8286.2005.00902.x. [DOI] [Google Scholar]
- Vasemägi A, Nilsson J, Primmer CR. Expressed sequence tag (EST) linked microsatellites as a source of gene associated polymorphisms for detecting signatures of divergent selection in Atlantic salmon (Salmo salar L.) Mol Biol Evol. 2005;22:1067–1076. doi: 10.1093/molbev/msi093. [DOI] [PubMed] [Google Scholar]
- Hancock JM. The contribution of slippage-like processes to genome evolution. J Mol Evol. 1995;41:1038–1047. doi: 10.1007/BF00173185. [DOI] [PubMed] [Google Scholar]
- Bachtrog D, Weiss S, Zangerl B, Brem G, Schlötterer C. Distribution of dinucleotide microsatellites in the Drosophila melanogaster genome. Mol Biol Evol. 1999;16:602–610. doi: 10.1093/oxfordjournals.molbev.a026142. [DOI] [PubMed] [Google Scholar]
- Katti MV, Ranjekar PK, Gupta VS. Differential distribution of simple sequence repeats in eukaryotic genome sequences. Mol Biol Evol. 2001;18:1161–1167. doi: 10.1093/oxfordjournals.molbev.a003903. [DOI] [PubMed] [Google Scholar]
- Molla M, Delcher A, Sunyaev S, Cantor C, Kasif S. Triplet repeat length bias and variation in the human transcriptome. Proc Natl Acad Sci USA. 2009;106:17095–17100. doi: 10.1073/pnas.0907112106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kantety RV, La Rota M, Matthews DE, Sorrells ME. Data mining for simple sequence repeats in expressed sequence tags from barley, maize, rice, sorghum and wheat. Plant Mol Biol. 2002;48:501–510. doi: 10.1023/A:1014875206165. [DOI] [PubMed] [Google Scholar]
- Bouza C, Hermida M, Millán A, Vilas R, Vera M, Fernández C, Calaza M, Pardo BG, Martínez P. Characterization of EST-derived microsatellites for gene mapping and evolutionary genomics in turbot. Anim Genet. 2008;39:666–670. doi: 10.1111/j.1365-2052.2008.01784.x. [DOI] [PubMed] [Google Scholar]
- Wang Y, Ren R, Yu Z. Bioinformatic mining of EST-SSR loci in Pacific oyster, Crassostrea gigas. Anim Genet. 2008;39:287–289. doi: 10.1111/j.1365-2052.2008.01701.x. [DOI] [PubMed] [Google Scholar]
- Stàgel A, Portis E, Toppino L, Rotino GL, Lanteri S. Gene-based microsatellite development for mapping and phylogeny studies in eggplant. BMC Genomics. 2008;9:357. doi: 10.1186/1471-2164-9-357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S, Zhang L, Matz M. Microsatellite characterization and marker development from public EST and WGS databases in the reef-building coral Acropora millepora (Cnidaria, Anthozoa, Scleractinia) J Hered. 2009;100:329–337. doi: 10.1093/jhered/esn100. [DOI] [PubMed] [Google Scholar]
- Areshchenkova T, Ganal MW. Comparative analysis of polymorphism and chromosomal location of tomato microsatellite markers isolated from different sources. Theor Appl Genet. 2002;104:229–235. doi: 10.1007/s00122-001-0775-2. [DOI] [PubMed] [Google Scholar]
- Primmer CR, Møller AP, Ellegren H. A wide-range survey of cross-species microsatellite amplification in birds. Mol Ecol. 1996;5:365–378. doi: 10.1111/j.1365-294x.1996.tb00327.x. [DOI] [PubMed] [Google Scholar]
- Galbusera P, van Dongen S, Matthysen E. Cross-species amplification of microsatellite primers in passerine birds. Conserv Genet. 2000;1:163–168. doi: 10.1023/A:1026587024065. [DOI] [Google Scholar]
- Wright TF, Johns PM, Walters JR, Lerner AP, Swallow JG, Wilkinson GS. Microsatellite variation among divergent populations of stalk-eyed flies, genus Cyrtodiopsis. Genet Res. 2004;84:27–40. doi: 10.1017/S0016672304006986. [DOI] [PubMed] [Google Scholar]
- Primmer CR, Painter JN, Koskinen MT, Palo JU, Merilä J. Factors affecting avian cross-species microsatellite amplification. J Avian Biol. 2005;36:348–360. doi: 10.1111/j.0908-8857.2005.03465.x. [DOI] [Google Scholar]
- FitzSimmons NN, Moritz C, Moore SS. Conservation and dynamics of microsatellite loci over 300 million years of marine turtle evolution. Mol Biol Evol. 1995;12:432–440. doi: 10.1093/oxfordjournals.molbev.a040218. [DOI] [PubMed] [Google Scholar]
- Rico C, Rico I, Hewitt G. 470 million years of conservation of microsatellite loci among fish species. Proc Biol Sci. 1996;263:549–557. doi: 10.1098/rspb.1996.0083. [DOI] [PubMed] [Google Scholar]
- Ezenwa VO, Peters JM, Zhu Y, Arevalo E, Hastings MD, Seppä P, Pedersen JS, Zacchi F, Queller DC, Strassmann JE. Ancient conservation of trinucleotide microsatellite loci in polistine wasps. Mol Phylogen Evol. 1998;10:168–177. doi: 10.1006/mpev.1998.0528. [DOI] [PubMed] [Google Scholar]
- Moore SS, Hale P, Bryne K. NCAM: a polymorphic microsatellite locus conserved across eutherian mammal species. Anim Genet. 1998;29:33–36. doi: 10.1046/j.1365-2052.1998.00234.x. [DOI] [PubMed] [Google Scholar]
- Webster MT, Smith NGC, Ellegren H. Microsatellite evolution inferred from human-chimpanzee genomic sequence alignments. Proc Natl Acad Sci USA. 2002;99:8748–8753. doi: 10.1073/pnas.122067599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vowles EJ, Amos W. Quantifying ascertainment bias and species-specific length differences in human and chimpanzee microsatellites using genome sequences. Mol Biol Evol. 2006;23:598–607. doi: 10.1093/molbev/msj065. [DOI] [PubMed] [Google Scholar]
- Kocher TD, Thomas WK, Meyer A, Edwards SV, Pääbo S, Villablanca FX, Wilson AC. Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers. Proc Natl Acad Sci USA. 1989;86:6196–6200. doi: 10.1073/pnas.86.16.6196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson JS. Fishes of the World. 4. New York: John Wiley and Sons, Inc; 2006. [Google Scholar]
- Ensembl. http://www.ensembl.org/index.html
- Shapiro MD, Marks ME, Peichel CL, Blackman BK, Nereng KS, Jonsson B, Schluter D, Kingsley DM. Genetic and developmental basis of evolutionary pelvic reduction in threespine sticklebacks. Nature. 2004;428:717–723. doi: 10.1038/nature02415. [DOI] [PubMed] [Google Scholar]
- Peichel CL, Ross JA, Matson CK, Dickson M, Grimwood J, Schmutz J, Myers RM, Mori S, Schluter D, Kingsley DM. The master sex-determination locus in threespine sticklebacks is on a nascent y chromosome. Curr Biol. 2004;14:1416–1424. doi: 10.1016/j.cub.2004.08.030. [DOI] [PubMed] [Google Scholar]
- Colosimo PF, Hosemann KE, Balabhadra S, Villarreal G Jr, Dickson M, Grimwood J, Schmutz J, Myers RM, Schluter D, Kingsley DM. Widespread parallel evolution in sticklebacks by repeated fixation of Ectodysplasin alleles. Science. 2005;307:1928–1933. doi: 10.1126/science.1107239. [DOI] [PubMed] [Google Scholar]
- Miller CT, Beleza S, Pollen AA, Schluter D, Kittles RA, Shriver MD, Kingsley DM. cis-regulatory changes in Kit ligand expression and parallel evolution of pigmentation in sticklebacks and humans. Cell. 2007;131:1179–1189. doi: 10.1016/j.cell.2007.10.055. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell MA, Foster SA. In: The Evolutionary Biology of the Threespine Stickleback. Bell MA, Foster SA, editor. Oxford: Oxford University Press; 1994. Introduction to the evolutionary biology of the threespine stickleback; pp. 1–27. [Google Scholar]
- Bell MA. In: The Evolutionary Biology of the Threespine Stickleback. Bell MA, Foster SA, editor. Oxford: Oxford University Press; 1994. Palaeobiology and evolution of threespine stickleback; pp. 438–471. [Google Scholar]
- Baker JA. In: The Evolutionary Biology of the Threespine Stickleback. Bell MA, Foster SA, editor. Oxford: Oxford University Press; 1994. Life history variation in female threespine stickleback; pp. 144–187. [Google Scholar]
- Chagné D, Chaumeil P, Ramboer A, Collada C, Guevara A, Cervera MT, Vendramin GG, Garcia V, Frigerio J-M, Echt C, Richardson T, Plomion C. Cross-species transferability and mapping of genomic and cDNA SSRs in pines. Theor Appl Genet. 2004;109:1204–1214. doi: 10.1007/s00122-004-1683-z. [DOI] [PubMed] [Google Scholar]
- Coulibaly I, Gharbi K, Danzmann RG, Yao J, Rexroad CE. Characterization and comparison of microsatellites derived from repeat-enriched libraries and expressed sequence tags. Anim Genet. 2005;36:309–315. doi: 10.1111/j.1365-2052.2005.01305.x. [DOI] [PubMed] [Google Scholar]
- Chabane K, Ablett GA, Cordeiro GM, Valkoun J, Henry RJ. EST versus genomic derived microsatellite markers for genotyping wild and cultivated barley. Genet Resour Crop Evol. 2005;52:903–909. doi: 10.1007/s10722-003-6112-7. [DOI] [Google Scholar]
- Fraser LG, McNeilage MA, Tsang GK, Harvey CF, De Silva HN. Cross-species amplification of microsatellite loci within the dioecious, polyploid genus Actinidia (Actinidiaceae) Theor Appl Genet. 2005;112:149–157. doi: 10.1007/s00122-005-0117-x. [DOI] [PubMed] [Google Scholar]
- Kondo H, Ino M, Hironori A, Ishizaki H, Iwami M. Multiple gene copies for bombyxin, an insulin-related peptide of the silkmoth Bombyx mori: structural signs for gene rearrangement and duplication responsible for generation of multiple molecular forms of bombyxin. J Mol Biol. 1996;259:926–937. doi: 10.1006/jmbi.1996.0370. [DOI] [PubMed] [Google Scholar]
- Kawahara R, Nishida M. Multiple occurrences of spiggin genes in sticklebacks. Gene. 2006;373:58–66. doi: 10.1016/j.gene.2006.01.008. [DOI] [PubMed] [Google Scholar]
- Hahn MW, Han MV, Han SG. Gene family evolution across 12 Drosophila genomes. PLoS Genet. 2007;3:3197. doi: 10.1371/journal.pgen.0030197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hashiguchi Y, Furuta Y, Kawahara R, Nishida M. Diversification and adaptive evolution of putative sweet taste receptors in threespine stickleback. Gene. 2007;396:170–179. doi: 10.1016/j.gene.2007.03.015. [DOI] [PubMed] [Google Scholar]
- Meguro Y, Takahashi H, Takeshima H, Nishida M, Goto A. Isolation and characterization of 13 microsatellite loci in the nine-spined stickleback (Pungitius pungitius) and cross-species amplification in 5 stickleback species (family Gasterosteidae) Conservation Genet Resour. 2009;1:31–34. doi: 10.1007/s12686-009-9007-x. [DOI] [Google Scholar]
- Koizumi N, Jinguji H, Takahashi H, Higuchi M, Takata K, Minezawa M, Takemura T, Mori A. Isolation and characterization of polymorphic microsatellite DNA markers in the Omono type of ninespine stickleback, genus Pungitius. Mol Ecol Notes. 2007;7:1315–1318. doi: 10.1111/j.1471-8286.2007.01867.x. [DOI] [Google Scholar]
- Takata K, Goto A, Yamazaki F. Genetic differences of Pungitius pungitius and P. sinensis in a small pond of the Omono River System. Jpn J Ichthyol. 1987;34:384–386. [Google Scholar]
- Schlötterer C, Amos W, Tautz D. Conservation of polymorphic simple sequence loci in cetacean species. Nature. 1991;354:63–65. doi: 10.1038/354063a0. [DOI] [PubMed] [Google Scholar]
- Endicott P, Ho SY, Metspalu M, Stringer C. Evaluating the mitochondrial timescale of human evolution. Trends Ecol Evol. 2009;24:515–521. doi: 10.1016/j.tree.2009.04.006. [DOI] [PubMed] [Google Scholar]
- Chakraborty R, Kimmel M, Stivers DN, Davison LJ, Deka R. Relative mutation rates at di-, tri-, and tetranucleotide microsatellite loci. Proc Natl Acad Sci USA. 1997;94:1041–1046. doi: 10.1073/pnas.94.3.1041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schug MD, Hutter CM, Wetterstrand KA, Gaudette MS, Mackay TF, Aquadro CF. The mutation rates of di-, tri- and tetranucleotide repeats in Drosophila melanogaster. Mol Biol Evol. 1998;15:1751–1760. doi: 10.1093/oxfordjournals.molbev.a025901. [DOI] [PubMed] [Google Scholar]
- Ellegren H, Moore S, Robinson N, Byrne K, Ward W, Sheldon BC. Microsatellite evolution - a reciprocal study of repeat lengths at homologous loci in cattle and sheep. Mol Biol Evol. 1997;14:854–860. doi: 10.1093/oxfordjournals.molbev.a025826. [DOI] [PubMed] [Google Scholar]
- Hutter CM, Schug MD, Aquadro CF. Microsatellite variation in Drosophila melanogaster and Drosophila simulans: a reciprocal test of the ascertainment bias hypothesis. Mol Biol Evol. 1998;15:1620–1636. doi: 10.1093/oxfordjournals.molbev.a025890. [DOI] [PubMed] [Google Scholar]
- Shikano T, Shimada Y, Herczeg G, Merilä J. History vs. habitat type: explaining the genetic structure of European nine-spined stickleback (Pungitius pungitius) populations. Mol Ecol. 2010;19:1147–1161. doi: 10.1111/j.1365-294X.2010.04553.x. [DOI] [PubMed] [Google Scholar]
- Scott KD, Eggler P, Seaton G, Rossetto M, Ablett EM, Lee LS, Henry RJ. Analysis of SSRs derived from grape ESTs. Theor Appl Genet. 2000;100:723–726. doi: 10.1007/s001220051344. [DOI] [Google Scholar]
- Thiel T, Michalek W, Varshney RK, Graner A. Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.) Theor Appl Genet. 2003;106:411–422. doi: 10.1007/s00122-002-1031-0. [DOI] [PubMed] [Google Scholar]
- Cho YG, Ishii T, Temnykh S, Chen X, Lipovich L, McCouch SR, Park WD, Ayer N, Cartinhour S. Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa) Theor Appl Genet. 2000;100:713–722. doi: 10.1007/s001220051343. [DOI] [Google Scholar]
- Li YC, Korol AB, Fahima T, Nevo E. Microsatellites within genes: structure, function, and evolution. Mol Biol Evol. 2004;21:991–1007. doi: 10.1093/molbev/msh073. [DOI] [PubMed] [Google Scholar]
- Fondon JW III, Garner HR. Molecular origins of rapid and continuous morphological evolution. Proc Natl Acad Sci USA. 2004;101:18058–18063. doi: 10.1073/pnas.0408118101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taggart JB, Hynes RA, Prodöhl PA, Ferguson A. A simplified protocol for routine total DNA isolation from salmonid fishes. J Fish Biol. 1992;40:963–965. doi: 10.1111/j.1095-8649.1992.tb02641.x. [DOI] [Google Scholar]
- Largiadèr CR, Fries V, Kobler B, Bakker TCM. Isolation and characterization of microsatellite loci from the three-spined stickleback (Gasterosteus aculeatus L.) Mol Ecol. 1999;8:342–344. [PubMed] [Google Scholar]
- Heckel G, Zbinden M, Mazzi D, Kohler A, Reckeweg G, Bakker TCM, Largiadèr CR. Microsatellite markers for the three-spined stickleback (Gasterosteus aculeatus L.) and their applicability in a freshwater and an anadromous population. Conserv Genet. 2002;3:79–81. doi: 10.1023/A:1014255027870. [DOI] [Google Scholar]
- Peichel CL, Nereng K, Ohgi KA, Cole BLE, Colosimo PF, Buerkle CA, Schluter D, Kingsley DM. The genetic architecture of divergence between threespine stickleback species. Nature. 2001;414:901–905. doi: 10.1038/414901a. [DOI] [PubMed] [Google Scholar]
- Colosimo PF, Peichel CL, Nereng K, Blackman BK, Shapiro MD, Schluter D, Kingsley DM. The genetic architecture of parallel armor plate reduction in threespine sticklebacks. PLoS Biol. 2004;2:635–641. doi: 10.1371/journal.pbio.0020109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mäkinen HS, Cano JM, Merilä J. Identifying footprints of directional and balancing selection in marine and freshwater three-spined stickleback (Gasterosteus aculeatus) populations. Mol Ecol. 2008;17:3565–3582. doi: 10.1111/j.1365-294X.2008.03714.x. [DOI] [PubMed] [Google Scholar]
- NCBI: National Center for Biotechnology Information. http://www.ncbi.nlm.nih.gov/genomes/leuks.cgi
- Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl Acids Symp Ser. 1999;41:95–98. [Google Scholar]
- Benson G. Tandem repeats finder: a program to analyze DNA sequences. Nucl Acids Res. 1999;27:573–580. doi: 10.1093/nar/27.2.573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, positions-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994;22:4673–4680. doi: 10.1093/nar/22.22.4673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- Martins WS, Lucas DCS, de Souza Neves KF, Bertioli DJ. WebSat - a web software for microsatellite marker development. Bioinformation. 2009;3:282–283. doi: 10.6026/97320630003282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brownstein MJ, Carpten JD, Smith JR. Modulation of non-templated nucleotide addition by Taq DNA polymerase: primer modifications that facilitate genotyping. BioTechniques. 1996;20:1004–1010. doi: 10.2144/96206st01. [DOI] [PubMed] [Google Scholar]
- Nei M. Molecular Evolutionary Genetics. New York: Columbia University Press; 1987. [Google Scholar]
- Goudet J. FSTAT (Version 1.2): a computer program to calculate F-statistics. J Hered. 1995;86:485–486. [Google Scholar]
- FSTAT 2.9.3. http://www2.unil.ch/popgen/softwares/fstat.htm
- Rice WR. Analyzing tables of statistical tests. Evolution. 1989;43:223–225. doi: 10.2307/2409177. [DOI] [PubMed] [Google Scholar]
- Van Oosterhout C, Hutchinson WF, Wills DP, Shipley P. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes. 2004;4:535–538. doi: 10.1111/j.1471-8286.2004.00684.x. [DOI] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Genetic variability of three-spined stickleback SSR markers in three populations of nine-spined sticklebacks.
List of three-spined stickleback SSR markers that were not polymorphic or successfully amplified in nine-spined sticklebacks.
Target genes and their function and location in the theree-spined stickleback genome, and PCR and sequencing primers for nine-spined sticklebacks.
Homology of nine-spined stickleback sequences in the three-spined stickleback genome and comparative genomic location of SSRs in these species.