Table 2.
Normalized distances and insertion costs for three comparisons.
| without missing homologs | missing homologs replaced | change | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Comparison | n | D | D/n | n | D | D/n | Δn | Δ D | 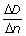 |
| Populusb -Vitis [7] | 2104 | 1092 | 0.52 | 6144 | 2545 | 0.41 | 4040 | 1453 | 0.36 |
| Populusb -Carica [2] | 2590 | 1461 | 0.56 | 7222 | 3466 | 0.48 | 4632 | 2005 | 0.43 |
| Ricinusc-Vitis | 13694 | 8283 | 0.60 | 18300 | 9931 | 0.54 | 4606 | 1682 | 0.37 |
| R.-V. correcteda | 13694 | 7715 | 0.56 | 18300 | 9365 | 0.51 | 4606 | 1684 | 0.37 |
a: contains corrections to distance due to scaffold fusions. b: Populus comparisons include rearrangements during the re-diploidization of the ancestral tetraploid. c: some missing homologs in the Ricinus comparisons are simply not sequenced, while in the other comparisons, missing homologs are virtually all absent from the genome.
