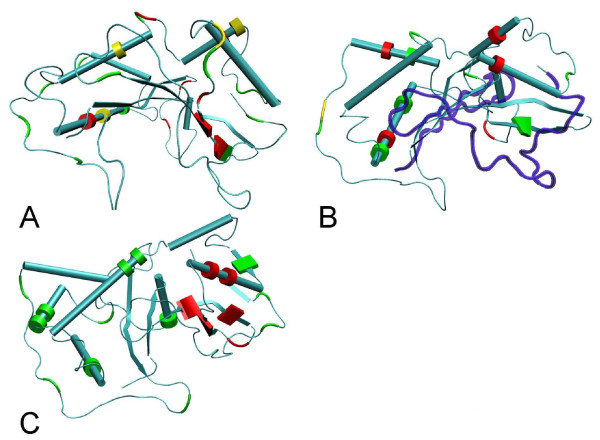
Figure 4.
Lysine amino acid residues mapped onto the S-RNase crystal structure. The Pyrus pyrifolia (Rosaceae; 1IQQ) structure is shown when using the Maloideae (Rosaceae) (A) and Prunus (Rosaceae) (B) datasets, while the Nicotiana alata (Solanaceae; 1IOO) structure is shown when using the Solanaceae dataset (C). Alpha helices are represented as tubes and beta-sheets as thin sheets. In panels A and B, lysine residues that are present in less than 50%, in between 50% and 75%, or in more than 75% frequency are colored in green, yellow and red, respectively. In panel C, the lysine residues shown to be important for ubiquitylation in Petunia inflata S3 RNase are labeled in red, while all other lysine residues are colored green. Regions marked in blue correspond to those regions of the S-RNase protein that were not inspected for the presence of lysine amino acid residues (see Material and Methods).