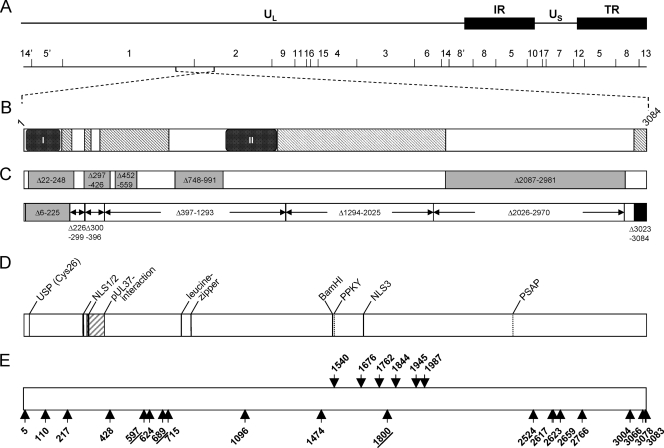
FIG. 1.
Schematic overview of PrV pUL36 and corresponding insertion mutants. (A) Diagram of the PrV genome with the unique long (UL) and unique short (US) regions as well as repeat regions (internal repeat, IR; terminal repeat, TR). The positions of BamHI restriction sites are indicated, and restriction fragments are numbered according to their size. (B) Schematic diagram of the UL36 open reading frame with conserved regions. Pfam analysis (4; http://www.sanger.ac.uk/Software/Pfam/) delineated two highly conserved PfamA domains within pUL36 homologs of herpesviruses of all three herpesvirus subfamilies [box I, Herpes_teg_N PrV (p)UL36, aa 11 to 178] and of alphaherpesviruses [box II, Herpes_UL36 PrV (p)UL36, aa 1000 to 1251] as well as PfamB domains (hatched rectangles) (6) (C) Known essential and nonessential regions in PrV pUL36. Nonessential regions are shown in gray, with the positions of the amino acids deleted in the corresponding constructs (6, 8). Deletions tested by Lee et al. (28) are shown below, marked by arrows. The essential C terminus is shown in black. Besides the N-terminal deletion Δ6-225, none of the truncated proteins was functional. (D) Predicted or identified motifs in pUL36: USP (Cys26), active-site cysteine of the deubiquitinating activity (24); pUL37 interaction domain (16, 27); NLS, nuclear localization signal (37); leucine zipper (27); and late domain motifs PPKY and PSAP (6). (E) Locations of linker insertions in pUL36 are indicated by arrows and the position of the amino acid immediately preceding the insertion. Insertions shown by arrows pointing upwards yielded functional proteins, while arrows pointing downwards indicate nonfunctional mutants. Insertions resulting in proteins which were impaired but not fully deficient in complementation are underlined. For orientation, the BamHI site separating BamHI fragments 1 and 2 is indicated.