Figure 2.
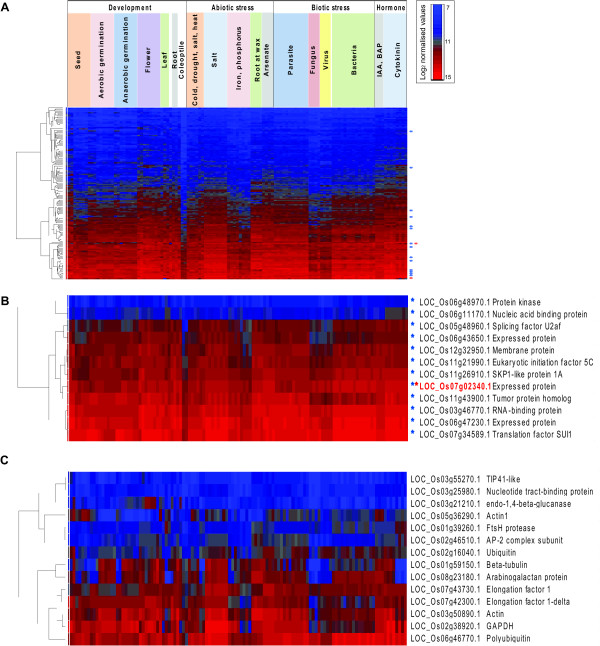
Analysis of stably expressed genes. A) Average linkage hierarchical clustering of the group of 151 probesets, based on CV criteria described in Figure 1. The genes are on the y-axis and the samples on the x-axis. The details of the treatments are outlined in Table 1. The scale is log2 normalised values where blue is low levels of transcript abundance and red is high levels of transcript abundance. Genes indicated by blue asterisk denotes novel reference genes indentified in this study, while red asterisk indicates genes previously defined as stably expressed in other studies [8,22]. B) The probesets indicated by blue asterisk (*) in A, were independently hierarchically clustered and analysed by QRT-PCR. C) Average linkage hierarchical clustering of the previously suggested/commonly used reference genes. The variation in transcript abundance across the various parameters is evident by the variation in colour intensity from left to right.