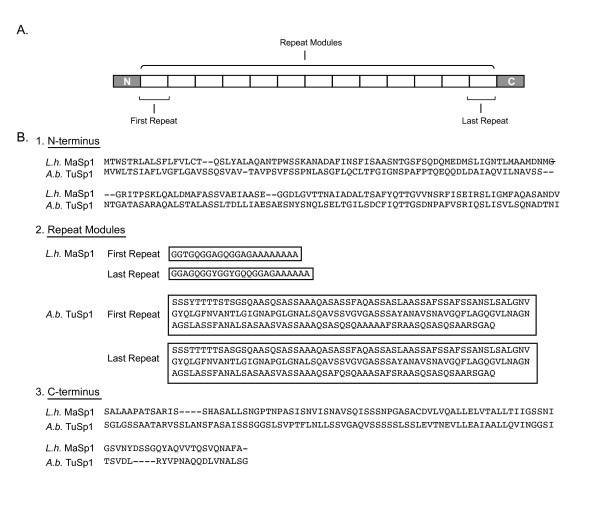
Figure 1.
Spidroin molecular organization and comparison of domain sequences from two paralogs. A. Schematic of spidroin primary structure showing short, non-repetitive N- and C-terminal domains flanking a long region of tandem sequence repeat modules. B. A comparison of full-length, divergent spidroin paralogs encoding the dragline silk protein MaSp1 from Latrodectus hesperus (L.h.) [Genbank: EF595246] and the egg-case silk protein TuSp1 from Argiope bruennichi (A.b.) [Genbank: AB242144], showing their (1) N-terminal domains, (2) the first and last repeat in each sequence and (3) their C-terminal domains. Dashes are alignment gaps. Note the varying repeat sequence length and composition between L.h. MaSp1 and A.b. TuSp1, in comparison to the high similarity across repeat modules within a spidroin.