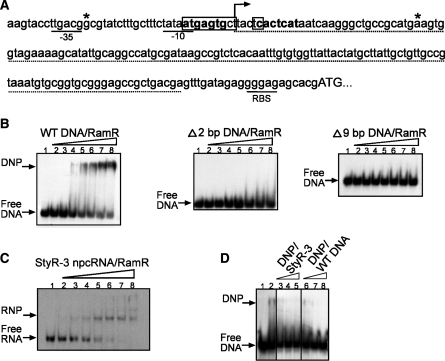
Figure 3.
Gel retardation analysis of the interaction of RamR protein with ramA gene regulatory region and StyR-3. (A) Sequence of ramA regulatory region. The −35, −10 and RBS sequences are underlined. Predicted RamR–DNA binding sites are indicated with bold letters. The 2-bp (Δ2 bp DNA) and 9-bp (Δ9 bp DNA) deletions within the ramA regulatory region are boxed. The transcriptional start site is indicated by an arrow. Asterisks indicate the 5′- and 3′-nt for the DNA fragment (WT DNA) used in the gel retardation assay. The StyR-3 sequence is indicated with a dotted line. (B) Gel retardation analysis of the interaction of RamR protein with the ramA gene regulatory region, showing wild-type DNA fragment (left), 2-bp (middle) and 9-bp (right) deletion fragments. (C) Interaction of RamR protein with StyR-3. (B and C) 32P-labeled DNA fragments or StyR-3 were incubated with S. typhi recombinant protein RamR as follows: lane 1, no protein added; (B and C) lanes 2–8: 50, 100, 150, 200, 400, 600 and 800 nM RamR protein was added, respectively; (D) Competition assay. 32P-labeled WT DNA was competed from RamR–DNP complex with increasing concentrations of unlabeled StyR-3 (lanes 3–5: 50, 250 and 500 nM, respectively) and WT DNA (lanes 6–8: 50, 250 and 500 nM, respectively). DNA: competitor molar ratios for lanes 3 and 6; 4 and 7; 5 and 8 were: 1:20; 1:100 and 1:200, respectively.