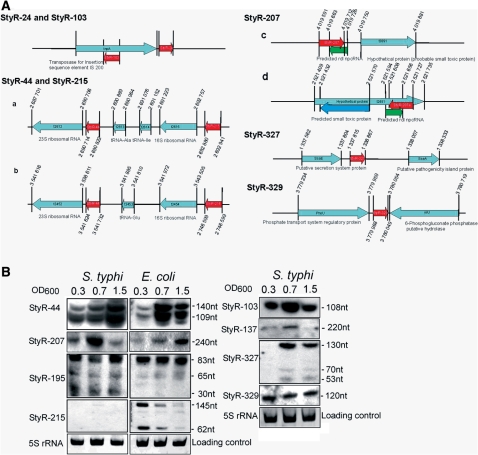
Figure 5.
Schematic representations and expression analyses of S. typhi repetitive npcRNA candidates. (A) Schematic representations of S. typhi repetitive npcRNA candidates. Coordinates of depicted genes are based on the completed genome of S. typhi Ty2 (AE014613). Locations of npcRNA candidates are depicted by red arrows. Known or predicted hypothetical genes are indicated with light blue arrows. Drawings are not to scale. StyR-24 and StyR-103 npcRNA candidates are each repeated 24 times in the S. typhi genome; only a schematic representation is shown. StyR-44 and StyR-215 are each repeated seven times in the S. typhi genome; only two rRNA operons are shown (a, b). StyR-207 and its schematic representation is depicted in (c); green arrow indicates putative S. typhi rdl npcRNA gene (annotated in this study). StyR-207a npcRNA (d) was predicted based on sequence similarity, putative ldr and rdl genes (annotated in this study) are depicted by blue and green arrows, respectively. (B) Northern blots showing expression patterns of repetitive npcRNA candidates at different growth stages of S. typhi and E. coli. OD600 values are indicated across the tops of the blots. OD600 0.3: lag phase-steady bacterial growth; OD600 0.7: log phase-exponential bacterial growth; and OD600 1.5: stationary growth phase. The npcRNA candidates and RNA lengths are indicated. As loading controls, a few examples of ethidium bromide-stained 5S rRNA are shown.