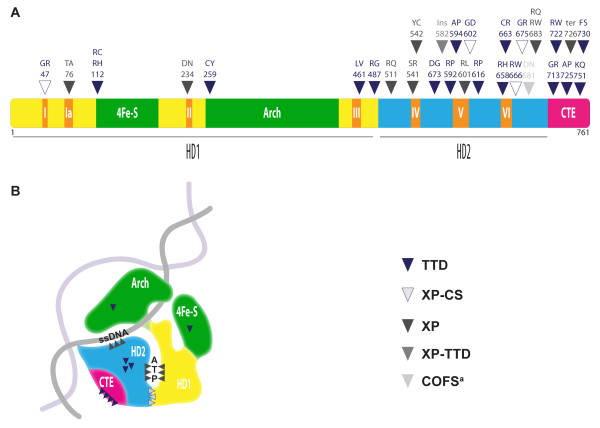
Figure 3.
(Adapted from [42]) (A) Linear aminoacid sequence of XPD, the positions of some of the (putative) causative mutations found in XPD syndromes are indicated [1,10,13,42,45,117]. Alterations considered to be completely inactivating [10] are not shown on this figure. With two exceptions (an aminoacid insertion at position 582 leading to XP-TTD and a non-sense mutation at position 726 leading to XP) all the mutations mapped in this drawing cause aminoacid substitutions. The phenotypic manifestation of these mutations is indicated by the color of the arrow as described in the legend. a COFS is caused by the allelic combination R616W (also found in TTD patients) and D681N (B) Cartoon representing the 3D structure of XPD according to the solved crystal structures of two bacterial homologues [42,45], the C terminus (pink) is absent in the bacterial proteins and therefore its location on the cartoon is not based on crystallography data [42]. Arrowheads represent schematically the predominant localization of mutations in XPD giving rise to the different disorders. XP mutations are found mainly in the vicinity of the helicase motifs on residues that form the channels binding either ATP or ssDNA. XP-CS mutations are at the interface between the two helicase domains and affect functional flexibility of the enzyme. In contrast TTD mutations appear to increase the flexibility of the XPD molecule affecting its framework stability and thereby also important intermolecular interactions such as those required to form the TFIIH complex. Additional TTD mutations are located in the C terminal part of the molecule outside of the catalytic core, Most of these mutations abolish interaction with the TFIIH subunit p44 [35].