FIG. 1.
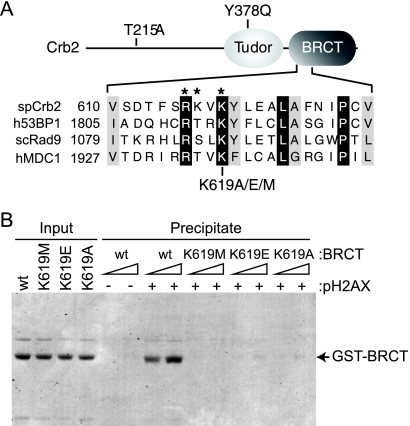
Crb2 pH2AX-binding mutations. (A) Top, schematic representation of Crb2 (not drawn to scale) with relevant mutations indicated. Bottom, protein sequence alignment of a portion of the BRCT phospho-binding motifs from Schizosaccharomyces pombe (sp) Crb2, human (h) 53BP1, human MDC1, and Saccharomyces cerevisiae (sc) Rad9. Identical residues are shaded black; similar residues are shaded gray. *, Crb2 phospho-binding residues. (B) The Crb2 BRCT domains specifically interact with pH2AX. Peptide pulldowns were performed as described in the text with C-terminal fission yeast H2A.1 peptides either unmodified or phosphorylated at Ser129 (see − or + pH2AX) and increasing amounts of the indicated recombinant Crb2 BRCT domain fragments (∼0.1 and 0.3 μM). After binding and washing, SDS-PAGE and Coomassie staining were used to visualize peptide-bound protein. A fraction of the total protein used for binding was also visualized (Input).