FIGURE 2.
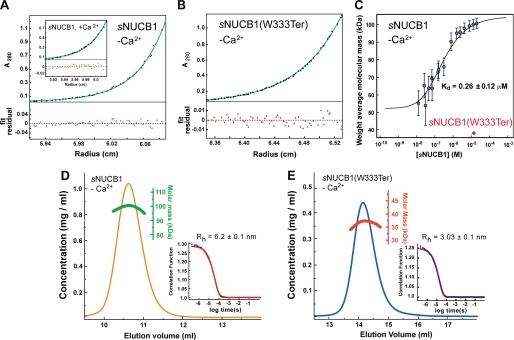
sNUCB1 forms a stable dimer mediated by its leucine zipper domain. A, we carried out AUC experiments to determine the oligomeric state of sNUCB1 in solution. The sedimentation equilibrium plot for Ca2+-free sNUCB1 (50 μm monomeric concentration) shows the evolution of a sample concentration curve resulting from the applied centrifugal force. The data were fit to a monomer-n-mer equilibrium model, which gave a measured molecular mass of 98.9 ± 0.41 kDa, consistent with a dimeric structure for sNUCB1 in solution (the theoretical molecular mass of monomeric sNUCB1 is ∼51 kDa). The inset shows the sedimentation equilibrium plot for Ca2+-bound sNUCB1, which gave essentially an identical molecular mass of 99.4 ± 0.78 kDa. B, AUC experiments were carried out on the truncation mutant, sNUCB1(W333Ter), which lacked the leucine zipper domain in the C-terminal region of the protein. The AUC sedimentation equilibrium plot for Ca2+-free sNUCB1(W333Ter) (70 μm monomeric concentration) gave a molecular mass of 35.2 ± 0.05 kDa, consistent with a monomeric species in solution (the theoretical molecular mass of sNUCB1(W333Ter) is ∼36.8 kDa). C, we studied the monomer-dimer equilibrium of sNUCB1 with MALS of fractions with increasing protein concentration collected from Superdex200 10/30 HR size-exclusion chromatography in 50 mm Tris-HCl, pH 8.0, 150 mm NaCl. The weight average molecular weight of the complex at a given protein concentration was determined from a nonlinear least square fit of a collection of values determined for the apex fractions of each eluting peak. A monomer-dimer association model of the values as a function of sNUCB1 concentration gave an apparent dissociation constant (Kd) for dimerization of 0.26 ± 0.12 μm. The error bars indicate the extent of variation in molecular mass determination originating from the light scattering measurement. We next measured DLS chromatograms for sNUCB1 (D) and sNUCB1(W333Ter) (E). Each protein sample was injected onto a Superdex200 10/30 HR column, and the refractive index detector was used to analyze the protein peak. The change in refractive index as a function of protein concentration was used to compute the molecular mass as shown in green for sNUCB1 and red for sNUCB1(W333Ter). The molecular mass corresponds to a dimer for sNUCB1 and a monomer for sNUCB1(W333Ter). The insets show the correlation functions as each protein diffuses through the solution in the DLS experiment. The diffusion coefficient from the correlation function gives the hydrodynamic radius (Rh) using Stokes equation. The computed Rh values were 6.2 and 3.03 nm for sNUCB1 and sNUCB1(W333Ter), respectively.