Table 1. Summary of crystal parameters and data-collection and refinement statistics for BT_3984 (PDB code 3cgh).
Values in parentheses are for the highest resolution shell.
| λ1 MADSe | λ2 MADSe | λ3 MADSe | |
|---|---|---|---|
| Space group | C2221 | ||
| Unit-cell parameters (Å) | a = 49.78, b = 125.27, c = 174.65 | ||
| Data collection | |||
| Wavelength (Å) | 0.9184 | 0.9791 | 0.9792 |
| Resolution range (Å) | 28.0–1.7 (1.76–1.70) | 28.1–1.7 (1.76–1.70) | 28.0–1.7 (1.76–1.70) |
| No. of observations | 221725 | 219734 | 219247 |
| No. of unique reflections | 60062 | 60017 | 59976 |
| Completeness (%) | 98.5 (97.1) | 98.1 (97.8) | 97.8 (97.3) |
| Mean I/σ(I) | 10.6 (2.5) | 10.1 (2.5) | 10.1 (2.5) |
| Rmerge on I† (%) | 5.6 (32.8) | 5.6 (32.9) | 5.5 (32.3) |
| Model and refinement statistics | |||
| Resolution range (Å) | 28.0–1.7 | ||
| No. of reflections (total) | 60025 | ||
| No. of reflections (test set) | 3037 | ||
| Completeness (%) | 99.2 | ||
| Data set used in refinement | λ1 MADSe | ||
| Cutoff criterion | |F| > 0 | ||
| Rcryst‡ | 0.140 | ||
| Rfree§ | 0.166 | ||
| Stereochemical parameters | |||
| Restraints (r.m.s. observed) | |||
| Bond angles (°) | 1.52 | ||
| Bond lengths (Å) | 0.016 | ||
| Average isotropic B value (Å2) | 18.1¶ | ||
| ESU†† based on Rfree (Å) | 0.083 | ||
| No. of protein residues | 507 | ||
| No. of protein atoms | 4035 | ||
| No. of waters | 704 | ||
| No. of other molecules | 2 (acetate, zinc) | ||
R
merge = 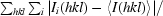
 .
.
R
cryst = 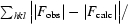
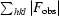 , where F
calc and F
obs are the calculated and observed structure-factor amplitudes, respectively.
, where F
calc and F
obs are the calculated and observed structure-factor amplitudes, respectively.
R free is the same as R cryst but for 5.1% of the total reflections that were chosen at random and omitted from refinement.
This value represents the total B that includes TLS and residual B components.
