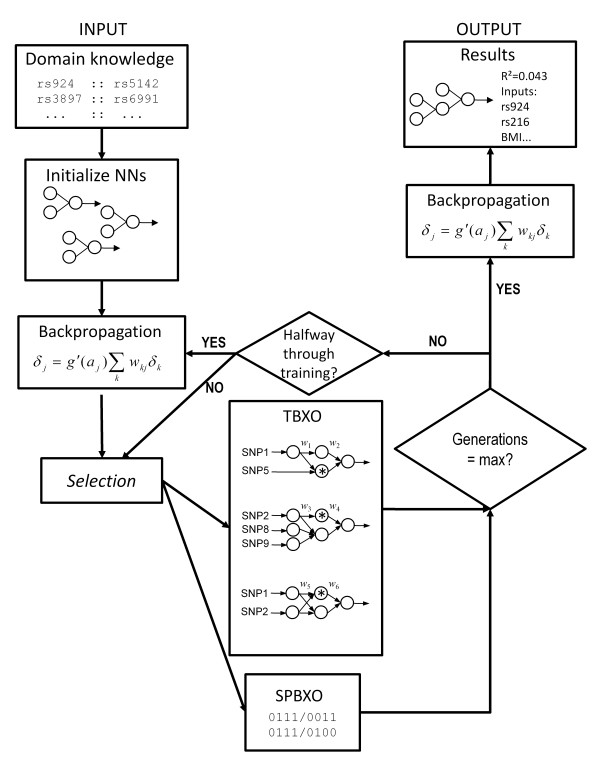
Figure 1.
ATHENA algorithm. The ATHENA algorithm begins by optionally accepting a list of SNP-SNP models which are derived from biological knowledge sources. This domain knowledge is used to initialize a proportion of the NN population. BP is used to optimize the initial weights. After a round of selection, GE is used to simultaneously optimize variable selection, NN architecture, and weights. Another round of BP takes place midway through training, and at the end of training. Crossover can occur via single point binary (SPBXO) or tree-based crossover (TBXO). In SPBXO, crossover occurs at the binary string level, but in TBXO, NNs are first translated, and crossover occurs at the binary genome level that results in a crossover at functionally similar root nodes. The NNs in this figure have either two or three inputs, corresponding to numerically coded values (-1, 0, 1) for SNP genotypes. A weight vector corresponds to each layer of weights in the NN. In TBXO, functionally analogous branches are crossed over, indicated by the asterisk, resulting in a 2-2-1 neural network with SNPs 1 and 2 as inputs. If SNPs 1 and 2 are the functional SNPs responsible for the gene-gene interaction and if the weight vectors on this NN are favorable, then this NN should be capable of modelling a gene-gene interaction between these two SNPs.